Professional Documents
Culture Documents
30 Tracing The Source of An Outbreak of Methicillin-Resistant
Uploaded by
vega_utukku1877Original Description:
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
30 Tracing The Source of An Outbreak of Methicillin-Resistant
Uploaded by
vega_utukku1877Copyright:
Available Formats
EPIDEMIOLOGY
Tracing the Source of an Outbreak of Methicillin-Resistant
Staphylococcus aureus in a Tertiary-Care Oncology
Hospital by Epidemiology and Molecular Methods
Patricia Cornejo-Jua rez,
1
Patricia Volkow-Ferna ndez,
1
Jose Sifuentes-Osornio,
2
Gabriela Echa niz-Aviles,
3
Adriana D az-Gonzalez,
1
Consuelo Vela zquez-Acosta,
1
Miriam Bobadilla-del-Valle,
2
Patricia Gordillo-Molina,
1
and Mar a E. Velazquez-Meza
3
This study describes the clinical and molecular characteristics of methicillin-resistant Staphylococcus aureus
(MRSA) isolates that emerged after an index case in a tertiary-care oncology hospital in Mexico City and
identies whether these isolates were related with the index case. All MRSA strains isolated from January 2006
until December 2007 were included. The clinical and demographic characteristics of patients were analyzed;
molecular typing by pulsed-eld gel electrophoresis was used to characterize the isolates. We included 44 MRSA
isolates from 55 patients. Thirty-eight patients (86.4%) were classied with nosocomial infection and the re-
mainder with healthcare-related infection. A single pulsed-eld gel electrophoresis pattern (C) was identied
with minor variations (two subtypes). The isolates analyzed were staphylococcal chromosome cassette mec type
II (related with the New York-Japan strain). This case underscores the need to intensify strategies that identify
and limit the spread of multiresistant pathogens imported by infected patients referred from other healthcare
centers.
Introduction
S
taphylococcus aureus is among the most important
microorganisms that causes human infections; it com-
prises part of human ora and is a major cause of nosocomial
infection. The spectrum of S. aureus infections includes toxic
shock syndrome, food poisoning, bloodstream infections,
endocarditis, osteomyelitis, meningitis, and dermatological
disorders ranging from minor infections and eczema to
blisters and scalded skin syndrome.
9
Methicillin-resistant S. aureus (MRSA) is a worldwide
problem associated with increased morbidity and mortality
among hospitalized patients; therefore, identication of the
risk factors inuencing its spread in hospitals is of great
concern for the prevention of nosocomial infections.
8
The
major reservoirs of staphylococci in hospitals are coloniza-
tion of patients and healthcare workers (HCW), as well as
infected patients. Carriers have a higher risk for developing
endogenous infection or transmitting infection to HCW and
patients. Transient hand carriage of the organism, usually by
HCW, account for the major mechanism for transmission.
9
Patients with cancer are placed at a high risk of developing
severe infections with Gram-positive organisms including
MRSA because of the frequent use of invasive devices, ex-
tensive surgeries, antineoplastic therapy, and immunosup-
pression associated with malignant disease.
5,17
Multiple DNA-based methods have been employed to
characterize isolates of MRSA that allow better discriminatory
power than phenotypic analyses.
23
These studies have dem-
onstrated the clonal spread of epidemic MRSA strains within
hospitals, between hospitals within a country, and even be-
tween countries and continents.
9,10,12
In Mexico, the preva-
lence of MRSA varies widely, ranging from 7% to 30%.
2,4
MRSA was rst isolated in July 2004; subsequently, it was
isolated sporadically. In January 2006, an outbreak was
identied; the index case was a female patient with osteo-
sarcoma with an exposed prosthesis infection who experi-
enced continuous secretion through the wound and a hospital
stay of 38 days over four different time periods. The case was
resolved nally with limb amputation during the patients
last hospitalization in March 2006. Since January 2006, the
month of the patients arrival at the hospital, MRSA was
1
Department of Infectious Diseases, Instituto Nacional de Cancerolog a, Mexico City, Mexico.
2
Laboratory of Clinical Microbiology, Instituto Nacional de Ciencias Medicas y Nutricio n Salvador Zubira n, Mexico City, Mexico.
3
Department of Vaccine Evaluation, Instituto Nacional de Salud Pu blica, Cuernavaca, Mexico.
MICROBIAL DRUG RESISTANCE
Volume 16, Number 3, 2010
Mary Ann Liebert, Inc.
DOI: 10.1089/mdr.2010.0048
203
continuously isolated fromdifferent patients with nosocomial
infection or healthcare-related infection (HCRI).
The purpose of the present study was to describe the
clinical and molecular characteristics of MRSA isolates
that emerged after the index case and to identify whether
these isolates were related with the index case.
Materials and Methods
Hospital setting
The Instituto Nacional de Cancerolog a (INCan) is a tertiary-
care oncology center, located in Mexico City; it has 150 beds
and a 6-bed intensive care unit. The hospital has 7,500 dis-
charges and 3,450 surgeries per year; 700 patients with long
indwelling central venous catheters are continuously cared
as outpatientsambulatory patients followed by the IV team
monthly, and 24,000 outpatient visits are performed for
chemotherapy-IV-infusion per year. The microbiology labora-
tory receives on average 14,000 samples annually. The hospital
has a Nosocomial Surveillance Programin operation since 1987.
Clinical data
The medical and laboratory records of patients with a
clinical isolate of MRSA during January 2006 to December
2007 were retrospectively reviewed. Only one isolate per case
of infection was included. Data were collected on demo-
graphic characteristics, underlying diseases, length of hospi-
talization, infection sites and complications, date of MRSA
isolation, type of specimen, susceptibility pattern, other
bacteria recovered from the same sample, antimicrobial
treatment, type and date of surgeries, use of central venous
catheter, and date of insertion. After MRSA isolation, we an-
alyzed the outcome (alive, death, or lost to follow-up) and
antimicrobial therapy for MRSA infection (type and dose of
antibiotics). Survival or follow-up time was calculated from
date of MRSA isolation to death, or to the date when the
infection was resolved. We considered antimicrobial therapy
to be appropriate if the antimicrobial agent prescribedshowed
clear inhibitory in vitro activity against the clinical isolate.
We screened for nasopharyngeal carriage of MRSA from
staff surgeons, medical and surgical residents, and nurses.
Antimicrobial susceptibility testing
All MRSA isolates were identied by standard microbio-
logical procedures.
15
Detection of methicillin resistance was
carried out according to the Clinical andLaboratory Standards
Institute guidelines using oxacillin disk test.
7
Antimicrobial
susceptibility testing was performed by the automated
MicroScan method (Dade-Behring, Sacramento, CA) for pen-
icillin, oxacillin, amikacin, amoxicillin, cefotaxime, cephalo-
thin, cefazolin, imipenem, trimethoprimsulfamethoxazole,
erythromycin, clarithromycin, clindamycin, ciprooxacin,
chloramphenicol, gentamicin, rifampin, tetracycline, and
vancomycin. S. aureus ATCC 29213 was utilized for quality
control as recommended by the Clinical and Laboratory
Standards Institute guidelines.
7
Molecular typing
Pulsed-eld gel electrophoresis (PFGE) was used to iden-
tify MRSA clones. Whole-genomic DNA was prepared as
described previously.
6
After digestion with SmaI endonu-
clease, DNA was separated in a CHEF-DRII apparatus
(Bio-Rad, Birmingham, United Kingdom).
6
Strains BK2464,
HD288, HU25, and EMRSA16, representing the NY/Japan-
USA, Pediatric-Portugal, Brazilian, and EMRSA-16-United
Kingdom clones, respectively, were included in PFGE gels
for comparison. These strains were kindly provided by Prof.
Herminia de Lencastre from the Molecular Genetic Labora-
tory, Instituto de Tecnologia Qu mica e Biologica da Uni-
versidade Nova de Lisboa. The criteria of Tenover et al. were
used to compare different clones.
22
Staphylococcal chromo-
some cassette mec (SCCmec) type was determined by a
multiplex polymerase chain reaction strategy. Strain BK2464
was used as SCCmec control.
19
Molecular analysis was per-
formed at the Instituto Nacional de Salud Pu blica in Cuer-
navaca, Mexico. Detection of mecA gene was performed at
the Instituto Nacional de Ciencias Medicas y Nutricio n in
Mexico City. Computer analysis of the banding patterns
obtained by PFGE was conducted with NTSYSpc ver.
2.0.2.11 software (Applied Biostatistics, Setauket, NY) after
visual inspection. Each gel included the reference strain
S. aureus NCTC 8325 to normalize PFGE proles. For cluster
analyses, Dice coefcients were calculated to compute matrix
similarity and were transformed into an agglomerative
cluster by the unweighted pair-group method with arith-
metic average.
21
Statistical methods
Comparison of categorical variables and percentages be-
tween groups was carried out by the Pearson chi-square test
or Fishers exact test, as appropriate. Logistic regression
analysis was performed to nd the association between
variables. A p-value of 0.05 was considered statistically
signicant. Statistical analysis was performed using Epi-Info
(ver. 6) and STATA (ver. 9.1) software.
Results
The rst case of this outbreak was a 23-year-old female
patient with a bone tumor diagnosed in August 2002 at an-
other hospital. The patient experienced a recurrence in
March 2005; the tumor was surgically removed and diag-
nosed as osteosarcoma. In October and November 2005, at
the same hospital, a tibial limb excision was performed and
a joint prosthesis was placed. Purulent secretion began to
drain a few days later and the prosthesis was exposed. The
patient received multiple antimicrobial treatments in addi-
tion to several surgical debridements. On January 2006, the
patient was referred to the INCan hospital, where osteomy-
elitis and joint prosthesis infection were diagnosed; the cul-
ture of the purulent secretion taken at hospital admission
yielded MRSA. Vancomycin and amikacin were initiated
and limb amputation was performed in March 2006 with
complete resolution of the infection.
From January 2006 to December 2007, 64 MRSA clinical
isolates were identied from 55 patients. Forty-four strains
(80%) were collected and stored at the laboratory of micro-
biology.
Thirty-eight infections (86.4%) were classied as nosoco-
mial, and the other six (13.6%) were HCRI. Over the study
period, the MRSA clinical isolates were recovered from di-
verse anatomical sites from different patients (Fig. 1).
204 CORNEJO-JUA
REZ ET AL.
Twenty-two patients were men (50%). Thirty-one patients
(70.5%) had a hematologic malignancy, 11 (25%) had a solid
tumor, and 2 (4.6%) had no neoplastic disease (biliary stenosis
of unknown cause and vertebral mycobacterial infection).
The most common infections were surgical site infection
(29.5%), pneumonia (27.3%), and bacteremia (13.6%). The
infection was polymicrobial in 13 patients (29.5%), and the
most common coinfectant microorganism was Escherichia coli
(n 7). Thirty-two patients (72.7%) had received antimicro-
bials during the previous 3 months. Other characteristics are
shown in Table 1.
Thirty-eight (86.4%) patients were diagnosed with an ep-
isode of nosocomial infection, of which 18 (47.4%) received
appropriate antimicrobial treatment: 6 of them died, 2 from
infection (mean time, 18.5 12 days) and 4 from another
cause (55.2 37.6 days) ( p 0.268). Twenty patients (45.5%)
did not receive appropriate antimicrobial treatment. Ten of
them died, six from infection (mean time, 3.7 2.3 days) and
four from another cause (mean time, 47.2 39.4 days)
( p 0.02). Mean survival for patients was 7.4 8.4 days. The
remaining patients recovered and were discharged.
There were six patients classied with HCRI; two of these
received appropriate treatment and were alive. Four patients
did not receive appropriate treatment and died: two from
infection and two from another cause. Mean survival time
was 51.2 35.9 days ( p 0.004).
We performed 173 nasopharyngeal cultures from staff and
found 24 (13.9%) methicillin-susceptible S. aureus and 2
MRSA (1.1%; 1 from a nurse and 1 from a surgical resident).
We were able to include only the latter isolate in the mo-
lecular analysis. However, the nurse did not receive treat-
ment, and a new nasal culture done at 16 weeks later did not
show MRSA. The surgical resident received ciprooxacin for
another cause, and a nasal culture taken at 8 weeks later was
negative.
All clinical isolates showed resistance to penicillin, oxacillin,
amoxicillin, cefotaxime, cephalothin, cefazolin, clindamycin,
imipenem, ciprooxacin, chloramphenicol, erythromycin, and
clarithromycin. No isolate was resistant to rifampin, amikacin,
tetracycline, gentamicin, trimethoprimsulfamethoxazole, or
vancomycin.
PFGE analysis identied a single clonal type, designated
pattern C, with two subtypes observed (C13 and C31) that
differed in up to three band positions. Lanes 1 and 10 rep-
resent lambda ladders used as molecular size (MW) markers;
lane 2, control strain NCTC 8325; lanes 3 and 4 (1 INCan
[index case] and 52 INCan), patterns C; lane 5 (4 INCan),
prole C13; lanes 69, control strains: BK2464 (New York/
FIG. 1. Date of isolation and type of sample from methicillin-resistant Staphylococcus aureus (MRSA) strains.
Table 1. Demographic and Clinical Characteristics
of Patients with Methicillin-Resistant
Staphylococcus aureus (n44)
Characteristic Patients with MRSA
Mean age (years) s.d. 46.1 18.4
Female sex, number (%) 22 (50)
Type of infection, number (%)
Surgical site infection 13 (29.5)
Pneumonia 12 (27.3)
Bacteremia 6 (13.6)
Bone-joint infections 5 (11.4)
Abscess 4 (9.1)
Other
a
4 (9.1)
Length of hospitalization s.d.
(range)
17.7 16 (279)
Patients who have received
antimicrobials, number (%)
32 (72.7)
Days of antimicrobials s.d.
(range)
15.1 13.7 (378)
Patients with recent surgery,
b
number (%)
30 (68.2)
Mean days from surgerys.d.
(range)
35.9 33.7 (1119)
Placement of CVC,
number (%)
30 (68.2)
Mean duration of CVC
placement s.d. (range)
89.3 127.3 (4540)
a
One endocarditis, one nephrostomy infection, one CVC site
infection, and one meningitis.
b
During the previous 3 months.
CVC, central venous catheter; MRSA, methicillin-resistant Staphy-
lococcus aureus; s.d., standard deviation.
OUTBREAK OF MRSA IN AN ONCOLOGY HOSPITAL IN MEXICO 205
Japan-USA clone), HD288 (Pediatric clone), HU25 (Brazilian
clone), and EMRSA-16 (United Kingdom clone) (Fig. 2).
Clone C showed a high degree of similarity with both the
Pediatric clone (89.5%) and New York/Japan clone (80.0%)
(Fig. 3). Characterization by SCCmec typing of clone C rep-
resentatives demonstrated that isolates in this clone were
SCCmec type II. Statistical analysis did not detect important
demographic or clinical differences in any of the three pa-
tients who presented clonal subtypes.
Discussion
In this report, we describe the rst MRSA outbreak in a
tertiary-care oncology hospital in Mexico City. Analysis of
MRSA isolates demonstrated that the source of this MRSA
outbreak was a transferred female patient with a compli-
cated bone-joint-prosthesis infection, who had been referred
from another hospital. Further, all 44 strains characterized
showed a multidrug-resistant prole, including b-lactams,
cephalosporins, carbapenems, macrolides, quinolones, clin-
damycin, and chloramphenicol.
We observed an increase in the rate of S. aureus during the
study period. MRSA is still an important cause of nosocomial
infections at the hospital, even with intensication of a pre-
ventive nosocomial infections campaign that includes rein-
forcement in hand washing policies, placement of alcohol
dispensers in all hospital and ambulatory-care areas, em-
phasizing of contact precautions for all patients with proven
MRSA infection, and active surveillance of cultures.
An outbreak described at an oncological center in Houston
identied 70 nosocomial MRSA isolates. There were 3 pre-
dominant clones in 25 of 33 strains (76%); 11 patients with
clone A (SCCmec type II). Following the implementation of
FIG. 2. Pulsed-eld gel electrophoresis proles of MRSA
clinical isolates from the Instituto Nacional de Cancerolog a
(INCan) and representatives of international MRSA clones.
FIG. 3. Dendrogram comparing MRSA clone C from INCan-Mexico with different international MRSA clones. For cluster
analyses, Dice coefcients were calculated to compute matrix similarity and transformed into an agglomerative cluster with
the unweighted pair-group method with arithmetic average.
206 CORNEJO-JUA
REZ ET AL.
interventional measures, a signicant decrease in nosocomial
MRSA isolates was noted.
13
It shows that measures im-
plemented by a good infection control program are an
additional tool to control bacterial resistance.
A study performed at a cancer hospital center in Ireland
reported an MRSA incidence of 20.3% in patients with febrile
neutropenia and bacteremia. A worrying trend was the high
level of methicillin resistance noted among S. aureus isolates
(89.3%), and the increasing rates in recent years.
14
Our study
showed the same trend.
Nevertheless, the causes of MRSA propagation remain
controversial, and contradictory conclusions have been
drawn from different studies.
20
Additionally, MRSA are also
resistant to different classes of antibiotics and have been
reported to acquire resistance to gentamicin and related
aminoglycosides.
9
All strains in this series demonstrated
susceptibility to gentamicin and glycopeptides.
It was found that 71% of the patients had received anti-
microbials recently. Detailed knowledge of susceptibility to
antimicrobial agents is essential to facilitate the development
of effective strategies to combat the growing problem of re-
sistance.
3
Clinicians should obtain material for culture and
susceptibility testing from all suspected sites including ab-
scesses and skin infections, especially those with necrotic
areas.
23
Molecular typing techniques allow identication of pan-
demic clones of MRSA and enable the monitoring of MRSA
clones circulating in different hospitals and at different time
intervals in a country.
12,18
The identication of well-dened
clonal groups provides a basis for understanding the dis-
semination of particular clones in the hospital environment
and will aid in preventing further dissemination of MRSA.
These techniques should help to predict the emergence of new
and even more serious strains of multidrug-resistant bacteria
(e.g., New York-Japan, Pediatric clone, USA 300, and USA
400). Such a rapid, extensive spread of MRSA can be due to
multiple episodes of horizontal transference and recombina-
tion of the mec gene, as suggested by clonal analysis.
10
Nosocomial-associated MRSA have been reported at
other hospitals in Mexico, with a wide geographic spread
of MRSA-specic clones in the country
11,24
similar to that
demonstrated with other clones in South America, Europe,
and the United States.
1,10
In this study, a predominant MRSA
clone (clone C) related with New York-Japan and Pediatric
clone was detected in this hospital outbreak; it showed three
PFGE subtypes and SCCmec type II. The MRSA clone found
in this outbreak has been already reported in Mexico at the
Hospital Civil Fray Antonio Alcalde in Guadalajara and at
the Hospital de Pediatr a del Centro Medico Nacional-Siglo
XXI in Mexico City, where it has been circulating since 1999
and 2001, respectively.
11,24
An outbreak described at an on-
cological center in Houston identied 70 nosocomial MRSA
isolates. There were three predominant clones in 25 of 33
strains (76%); 11 patients with clone A(SCCmec type II).
13
It is
noteworthy to mention that the rst reports of vancomycin-
resistant MRSA were from genetic linage USA 100, which is
also known as the New York-Tokyo clone, which is very
similar to the Mexican clone C. The SCCmec typing system is
used to distinguish between healthcare-associated MRSA
and community-acquired MRSA clones.
25
In the United
States, two clones, designated as USA 300 and USA 400 by
the Centers for Disease Control and Prevention, have been
identied as the primary clone types that cause community-
acquired MRSA infections. These clones have frequently
been associated with the Panton-Valentine leukocidin viru-
lence factor and the presence of SCCmec type IV.
16
In contrast,
hospital-acquired or healthcare-associated MRSA strains
usually lack genes for Panton-Valentin leukocidin and are
associated with other SCCmec (types I, II, or III). MRSA with
SCCmec type II tend to be resistant to b-lactams, carbapenems
aminoglycosides, macrolides, clindamycin, uoroquinolones,
and glycopeptides, whereas SCCmec type IV are usually re-
sistant to b-lactams and erythromycin but retain susceptibility
to clindamycin, trimethroprimsulfamethoxazole, and uor-
oquinolones. Healthcare-associated genotypes are frequently
multidrug-resistant.
14,16
This study highlights the need for hospital clinicians to be
aware of the common bacteria isolated in their unit and their
unusual antibiotic susceptibility. Microbes possess the ca-
pacity to evolve in response to their environment; the major
impetus for developing resistance is selective pressure re-
sulting from antibiotic use.
3,9
Although MRSA is now en-
demic at the hospital, the number of MRSA isolates identied
in the past 18 months has decreased since infection prevention
and control measures have been implemented.
13
Identication of carriers, isolation of colonized or infected
patients, use of barrier precautions, and reinforcement of
health workers hand washing are important strategies
for detection and limiting MRSA spread. In the setting of
clinical bacteremia, the clinician should immediately initiate
empirical therapy with appropriate antimicrobials to cover
the possibility of MRSA. Until isolation conrms or denies
this, infectious disease consultation is warranted in these
cases.
In conclusion, our data show that MRSA has spread to the
entire hospital from an index patient who arrived with a
surgical site infection from another institution. This case
underscores the need to intensify strategies that identify and
limit the spread of multiresistant pathogens in tertiary-care
hospitals by infected patients referred from other healthcare
centers.
Disclosure Statement
All authors report no conicts of interest relevant to this
article.
References
1. Aires de Sousa, M., M. Miragaia, I.S. Sanches, A. Avila, I.
Adamson, S.T. Casagrande, M.C. Brandileone, R. Palacio,
L. DellAcqua, M. Hortal, T. Camou, A. Rossi, M.M.E.
Velazquez, A.G. Echaniz, S.F. Solo rzano, I. Heitmann, and
H. de Lencastre. 2001. Three-year assessment of methicillin-
resistant Staphylococcus aureus clones in Latin America from
1996 to 1998. J. Clin. Microbiol. 39:21972205.
2. Alpuche, A.C., F.C. Avila, D.M. Espinoza, B.D. Go mez, and
J.I. Santos. 1989. Perles de sensibilidad antimicrobiana de
Staphylococcus aureus en un hospital pedia trico: prevalencia
de resistencia a meticilina. Bol. Med. Hosp. Infant. Mex.
11:697699.
3. Baddour, M.M., M.M. Abuelkheir, and A.J. Fatani. 2006.
Trends in antibiotic susceptibility patterns and epidemi-
ology of MRSA isolates from several hospitals in Riyadh,
Saudi Arabia. Ann. Clin. Microbiol. Antimicrob. 5:30.
OUTBREAK OF MRSA IN AN ONCOLOGY HOSPITAL IN MEXICO 207
4. Caldero n, J.E., and B.R. Espinoza Avila. 2002. Epidemiol-
ogy of drug resistance: the case of Staphylococcus aureus and
coagulase negative staphylococci infections. Salud Publica
Mex. 44:108112.
5. Celkan, T., A. Ozkan, H. Apak, S. Diren, G. Can, L. Yuksel,
and I. Yildiz. 2002. Bacteremia in childhood cancer. J. Trop.
Pediatr. 48:373377.
6. Chung, M., H. de Lencastre, P. Matthews, A. Tomasz, I.
Adamsson, M. Aires de Sousa, T. Camou, C. Cocuzza,
A. Corso, I. Couto, A. Domnguez, M. Gniadkowski, R.
Goering, A. Gomes, K. Kikuchi, A. Marchese, R. Mato, O.
Melter, D. Oliveira, R. Palacio, R. Sa-Leao, I. Santos San-
ches, J.H. Song, P.T. Tassios, and P. Villari. 2000. Molecular
typing of methicillin-resistant Staphylococcus aureus by
pulsed-eld gel electrophoresis: comparison of results ob-
tained in a multilaboratory effort using identical protocols
and MRSA strains. Microb. Drug Resist. 6:189198.
7. [CLSI] Clinical and Laboratory Standards Institute. 2006.
Performance Standards for Antimicrobial Susceptibility
Testing. 13th Informational Supplement, Vol. 25, no. 26.
Approved Standard M2-A7. CLSI, Vayne, PA.
8. Cosgrove, S.E., G. Sakoulas, E.N. Perencevich, M.J.
Schwaber, A.W. Karchmer, and Y. Carmeli. 2003. Com-
parison of mortality associated with methicillin-resistant and
methicillin-susceptible Staphylococcus aureus bacteraemia: a
meta-analysis. Clin. Infect. Dis. 36:5359.
9. Dar, J.A., M.A. Thoker, J.A. Khan, A. Ali, M.A. Khan, M.
Rizwan, K.H. Bhat, M.J. Dar, N. Ahmed, and S. Ahmad.
2006. Molecular epidemiology of clinical and carrier strains
of methicillin resistant Staphylococcus aureus (MRSA) in the
hospital settings of north India. Ann. Clin. Microbiol. Anti-
microb. 5:22.
10. Da Silva Coimbra, M.V., M.C. Silva-Carvalho, H. Wis-
plinghoff, G.O. Hall, S. Tallent, S. Wallance, M.B. Ed-
mond, A.M. Figueiredo, and R.P. Wenzel. 2003. Clonal
spread of methicillin-resistant Staphylococcus aureus in a
large geographic area of the United States. J. Hosp. Infect.
53:103110.
11. Echaniz, A.G., M.E. Velazquez, M. Aires de Sousa, O.R.
Morfn, N.E. Rodrguez, B.N. Carnalla, A.S. Esparza, and
H. de Lencastre. 2006. Molecular characterization of a
dominant methicillin-resistant Staphylococcus aureus (MRSA)
clone in a Mexican hospital (19992003). Clin. Microbiol.
Infect. 12:1228.
12. Faria, N.A., J.A. Carrinco, D.C. Oliveira, M. Ram rez, and
H. de Lencastre. 2008. Analysis of typing methods for epi-
demiological surveillance of both methicillin-resistant and
methicillin-susceptible Staphylococcus aureus strains. J. Clin.
Microbiol. 46:136144.
13. Ghanem, G., R.Y. Hachem, R.F. Chemaly, T. Dvorak, K.
Hulten, L. Graviss, and I.I. Raad. 2008. The role of molec-
ular methods in the prevention of nosocomial methicillin-
resistant Staphylococcus aureus clusters in cancer patients.
Am. J. Infect. Control 36:656660.
14. Graham, P.L., III, S.X. Lin, and E.L. Larson. 2006. A U.S.
population-based survey of Staphylococcus aureus coloniza-
tion. Ann. Intern. Med. 144:318325.
15. Ji, Y. 2007. Methicillin-resistant Staphylococcus aureus
(MRSA) protocols. In Y. Ji (ed.), Methods in Molecular
Biology. Humana Press, New Jersey, pp. 227258.
16. Mollering, R.C. 2006. The growing menace of community-
acquired methicillin-resistant Staphylococcus aureus. Ann.
Intern. Med. 144:368370.
17. Morris, P.G., T. Hassan, M. McNamara, A. Hassan, R.
Wiig, L. Grogan, O. Breathnach, E. Smyth, and H. Hum-
phreys. 2008. Emergence of MRSA in positive blood cultures
from patients with febrile neutropeniaa cause for concern.
Support. Care Cancer 16:10851088.
18. Oliveira, D.C., A. Tomasz, and H. de Lencastre. 2002. Se-
crets of success of a human pathogen: molecular evolution
of pandemic clones of methicillin-resistant Staphylococcus
aureus. Lancet Infect. Dis. 2:180189.
19. Oliveira, D.C., and H. de Lencastre. 2002. Multiplex PCR
strategy for rapid identication of structural types and
variants of the mec element in methicillin-resistant Staphy-
lococcus aureus. Antimicrob. Agents Chemother. 46:2155
2161.
20. Riley, L.W. 2004. Hospital infections: Staphylococcus aureus.
In L.W. Riley (ed.), Molecular Epidemiology of Infectious
Diseases: Principles and Practices. ASM Press, Washington,
DC, pp. 249280.
21. Struelens, M.J., A. Deplano, C. Godard, N. Maes, and
E. Serruys. 1992. Epidemiological typing and delineation of
genetic relatedness of methicillin resistant Staphylococcus
aureus by macro-restriction analysis of genomic DNA by
using pulsed eld electrophoresis. J. Clin. Microbiol.
30:25992605.
22. Tenover, F.C., R.D. Arbeit, R.V. Goering, P.A. Mickelsen,
B.E. Murray, D.H. Persing, and B. Swaminathan. 1995.
Interpreting chromosomal DNA restriction patterns pro-
duced by pulsed-eld gel electrophoresis: criteria for bacte-
rial strain typing. J. Clin. Microbiol. 33:22332239.
23. van Belkum, A., M. Struelens, A. de Visser, H. Verbrugh,
and M. Tibayrenc. 2001. Role of genomic typing in taxon-
omy, evolutionary genetics, and microbial epidemiology.
Clin. Microbial. Rev. 14:547560.
24. Velazquez, M.M.E., M. Aires de Sousa, A.G. Echaniz,
S.F. Solo rzano, N.G. Miranda, S.J. Silva and H. de Len-
castre. 2004. Surveillance of methicillin-resistant S. aureus in
a pediatric hospital in Mexico City during a 7-year period
(1997 to 2003): clonal evolution and impact of infection
control. J. Clin. Microbiol. 42:38773880.
25. Zhang, K.J.A., A. McClure, S. Elsayed, T. Louie, and
J.M. Conly. 2005. Novel multiplex PCR assay for charac-
terization and concomitant subtyping of staphylococcal
cassette chromosome mec types I to V in methicillin-resistant
Staphylococcus aureus. J. Clin. Microbiol. 43:50265033.
Address correspondence to:
Mar a E. Velazquez-Meza, Ph.D.
Department of Vaccine Evaluation
Instituto Nacional de Salud Pu blica
Universidad 655, Santa Mar a Ahuacatitlan,
Cerrada de los Pinos y Caminera
Cuernavaca 62100
Morelos
Mexico
E-mail: mevelaz@insp.mx
208 CORNEJO-JUA
REZ ET AL.
You might also like
- Herpesvirus Infections of The Nervous SystemDocument21 pagesHerpesvirus Infections of The Nervous SystemMarcela Garzon O VelezNo ratings yet
- Pathophysiology, Transmission, Diagnosis, and Treatment of Coronavirus Disease 2019 (COVID-19) A ReviewDocument13 pagesPathophysiology, Transmission, Diagnosis, and Treatment of Coronavirus Disease 2019 (COVID-19) A ReviewJose de Jesus Ayala LeyvaNo ratings yet
- Cortical Factin Stabilization Generates Apical LateralDocument25 pagesCortical Factin Stabilization Generates Apical Lateralvega_utukku1877No ratings yet
- Patogenicidad y VirulenciaDocument11 pagesPatogenicidad y Virulenciavega_utukku1877No ratings yet
- 30 Tracing The Source of An Outbreak of Methicillin-ResistantDocument6 pages30 Tracing The Source of An Outbreak of Methicillin-Resistantvega_utukku1877No ratings yet
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (895)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (588)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (266)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2259)
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (120)
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- CCNA Training New CCNA - RSTPDocument7 pagesCCNA Training New CCNA - RSTPokotete evidenceNo ratings yet
- Anderson, Poul - Flandry 02 - A Circus of HellsDocument110 pagesAnderson, Poul - Flandry 02 - A Circus of Hellsgosai83No ratings yet
- Bardonna MenuDocument16 pagesBardonna MenuFarley ElliottNo ratings yet
- GB GW01 14 04 02Document2 pagesGB GW01 14 04 02Muhammad LukmanNo ratings yet
- 23001864Document15 pages23001864vinodsrawat33.asiNo ratings yet
- Solar Charge Controller: Solar Car Solar Home Solar Backpack Solar Boat Solar Street Light Solar Power GeneratorDocument4 pagesSolar Charge Controller: Solar Car Solar Home Solar Backpack Solar Boat Solar Street Light Solar Power Generatorluis fernandoNo ratings yet
- Recruitment and Selection in Canada 7Th by Catano Wiesner Full ChapterDocument22 pagesRecruitment and Selection in Canada 7Th by Catano Wiesner Full Chaptermary.jauregui841100% (51)
- F24 60manual (New)Document14 pagesF24 60manual (New)Robert CumpaNo ratings yet
- Book Index The Art of Heavy TransportDocument6 pagesBook Index The Art of Heavy TransportHermon Pakpahan50% (2)
- Coffee Quality Manual by Abra Rand Nig Use IDocument25 pagesCoffee Quality Manual by Abra Rand Nig Use IIpungNo ratings yet
- Water Filling MachineDocument15 pagesWater Filling Machinepallab D RozarioNo ratings yet
- MSDS DowthermDocument4 pagesMSDS DowthermfebriantabbyNo ratings yet
- Bulk Material/Part Ppap Process Checklist / Approval: Required?Document32 pagesBulk Material/Part Ppap Process Checklist / Approval: Required?krds chidNo ratings yet
- Clocks (New) PDFDocument5 pagesClocks (New) PDFAbhay DabhadeNo ratings yet
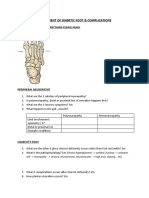
- Assessment of Diabetic FootDocument7 pagesAssessment of Diabetic FootChathiya Banu KrishenanNo ratings yet
- 988611457NK448908 Vehicle Scan ReportDocument5 pages988611457NK448908 Vehicle Scan ReportVictor Daniel Piñeros ZubietaNo ratings yet
- Psle Science Keywords !Document12 pagesPsle Science Keywords !Aftertea CarousellNo ratings yet
- Ict 2120 Animation NC Ii Week 11 20 by Francis Isaac 1Document14 pagesIct 2120 Animation NC Ii Week 11 20 by Francis Isaac 1Chiropractic Marketing NowNo ratings yet
- 500 TransDocument5 pages500 TransRodney WellsNo ratings yet
- Terminals of Ecm: E3 E4 E5 E6Document2 pagesTerminals of Ecm: E3 E4 E5 E6jeremih alhegn100% (1)
- MMW ReviewerDocument3 pagesMMW ReviewerMarcSaloj NeryNo ratings yet
- Kinder DLL Week 8Document15 pagesKinder DLL Week 8Jainab Pula SaiyadiNo ratings yet
- Medical GeneticsDocument4 pagesMedical GeneticsCpopNo ratings yet
- Matters Signified by The Sublord of 11th Cusp in KP SystemDocument2 pagesMatters Signified by The Sublord of 11th Cusp in KP SystemHarry HartNo ratings yet
- LinkageDocument9 pagesLinkageHarshu JunghareNo ratings yet
- FebvreDocument449 pagesFebvreIan Pereira AlvesNo ratings yet
- Quartile1 PDFDocument2 pagesQuartile1 PDFHanifah Edres DalumaNo ratings yet
- Rachel Joyce - A Snow Garden and Other Stories PDFDocument118 pagesRachel Joyce - A Snow Garden and Other Stories PDFИгорь ЯковлевNo ratings yet
- Arts Class: Lesson 01Document24 pagesArts Class: Lesson 01Lianne BryNo ratings yet
- Reading Part 2Document14 pagesReading Part 2drama channelNo ratings yet