Professional Documents
Culture Documents
The solution of crystallization problem was introduced around twenty years ago, with the introduction of crystallization screening methods. Here reported some of the factors which affect protein crystallization, solubility, Concentration of precipitant, concentration of macromolecule, ionic strength, pH, temperature, and organism source of macromolecules, reducing or oxidizing environment, additives, ligands, presence of substrates, inhibitors, coenzymes, metal ions and rate of equilibration. The aim of this paper to give very helpful advice for crystallization.
Uploaded by
Peertechz Publications Inc.Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
The solution of crystallization problem was introduced around twenty years ago, with the introduction of crystallization screening methods. Here reported some of the factors which affect protein crystallization, solubility, Concentration of precipitant, concentration of macromolecule, ionic strength, pH, temperature, and organism source of macromolecules, reducing or oxidizing environment, additives, ligands, presence of substrates, inhibitors, coenzymes, metal ions and rate of equilibration. The aim of this paper to give very helpful advice for crystallization.
Uploaded by
Peertechz Publications Inc.Copyright:
Available Formats
Global Journal of Biotechnology and Biomaterial Science
Mohnad Abdalla1*, Wafa Ali Eltayb1,
Abdus Samad1, Elshareef SHM1 and
TIM Dafaalla2
Hefei National Laboratory for Physical Sciences at the
Microscale and School of Life Sciences, University of Science
and Technology of China, Hefei, Anhui 230027, Peoples
Republic of China
College of Plant Science, Jilin University, Changchun
130062, China
Mini Review
Important Factors Influencing
Protein Crystallization
Dates: Received: 16 September, 2016; Accepted:
29 September, 2016; Published: 30 September,
2016
*Corresponding author: Mohnad Abdalla,
Hefei National Laboratory for Physical Sciences
at the Microscale and School of Life Sciences,
University of Science and Technology of
China, Hefei, Anhui 230027, PR China, E-mail:
Abstract
The solution of crystallization problem was introduced around twenty years ago, with the
introduction of crystallization screening methods. Here reported some of the factors which affect
protein crystallization, solubility, Concentration of precipitant, concentration of macromolecule, ionic
strength, pH, temperature, and organism source of macromolecules, reducing or oxidizing environment,
additives, ligands, presence of substrates, inhibitors, coenzymes, metal ions and rate of equilibration.
The aim of this paper to give very helpful advice for crystallization.
www.peertechz.com
Keywords: Proteins; Crystallization; Isoelectric point;
pH; Solubility; Stability
Introduction
X-ray crystallography has provided 3D structures of thousands
of proteins. In spite of these advances, many factors continue to
be problem that can lead to unsuccessful proteins crystallization.
We always know theoretical pI, molecular weight and amino-acid
composition, while pH and salt concentration are some of the
variables that can be expected from other similar known structure.
Yet, a protein behavior depends very much on the environment it
is in.
Proteins are generally present in a biological sample as their
native state. They are very often associated with other proteins
and integrated into large complexes. In most cases they are not
soluble in their innate state after isolation as result of that must be
denatured to help solubilization and Crystallization. Initial protein
crystal commonly need to screen conditions using little as possible to
improve the optimization methods for protein crystallization.
Crystallization is thermodynamically process. Once begin, will
continue under kinetic control until super saturation is lost. The
crystallization process affected by physical conditions of the solution,
solution solubility, the presence of impurities, nucleation, solution
saturation and degree of super saturation, crystal growth, including
solution composition, pH and temperature, and to date is not fully
understood.
Prediction of secondary structure, solubility, domain organization,
stability, signal peptide, hydrophobicity and pH, can provide useful
information for crystallization strategies and salt prediction tools
it need to be develop. in general proteins has low molecular weight
easier for crystallization than high molecular weight, single-domain
easier for crystallization than multi-domain and an oligomeric state
multimer more likely to crystallize than a monomer high molecular
weight.
Buffer is most straight forward way to make a crystallization
problem by effect the protein behave. There are many rules and
different protein crystallization methods, some of it has been
developed during the recent years. However, Protein crystallization
still represents a great challenge for bio crystallography. The
cumulation of crystallization information in crystallization databases
and in structural articles it allow us to design crystallization
experiments depending on the character of the protein. Because
there is no ideal procedure for crystallization, our article discusses
the crystallization problem try to simplify the composition of the
crystallization and give some solution advice.
Protein solubility
Protein solubility is a common complex interaction problem
between the physiochemical nature of the proteins, rate of protein
synthesis, amino acid composition, the Protein concentration, type
of salt present in the buffer, the concentration of the salt used, ionic
strength, pH, temperature, osmotic pressure, cellular location of
expression, and cellular tools or chaperones are all important in
protein solubility. Protein solubility may decrease in the presence of
complexes with lipids, nucleic acids or other nonprotein.
It is important to select better option for tag in the target proteins.
Unfortunately, there is no consolidated system for rapid screening to
differentiate the best tag.
Addition L-Arg and L-Glu at 50 mM to the buffer can increase
solubility up to 8.7 times, because they can in preventing protein
aggregation and precipitation as well as increase the long-term
stability also it did not prevent specific protein-protein and proteinRNA interactions [1].
Normally when there is solubility issues, the first thing we need
to check it the buffer composition, varying pH and salt strength it can
make a big difference to solubility. Then try solubilising agents, e.g
NDSBs or detergents. Protein solubility can improved by selecting
only a soluble domain for expression, and delete the hydrophobic
domain. Moreover there are solubility prediction tools in the internet
it can be useful.
In order to be sure target protein is soluble culture need to be
lysate (lysozime 1mg/ml on ice 1hour, it should be under non
Citation: Abdalla M, Ali Eltayb W, Samad A, Elshareef SHM, Dafaalla TIM (2016) Important Factors Influencing Protein Crystallization. Glob J Biotechnol
Biomater Sci 2(1): 025-028.
025
Abdalla et al. (2016)
denaturation condition) and centrifugation at 35000 x g. then use an
aliquot of supernatant for SDS-PAGE. In case of protein is soluble
well found in the supernatant.
Detergents
Detergents are commonly used at concentrations of 14%. It
useful in solubilizing by disrupt hydrophobic interactions between
and within proteins. The detergent used for solubilization does not
requisite to be the same for purification and crystallization, also
the presence of various detergents perhaps inhibit the activity of
different proteases. When despite all efforts not work then it needs
denaturation and solubilization of the target protein. This is done by
a denaturing agent e.g. guanidine or urea under reducing conditions
(~20 mM DTT). A western blot necessary for the supernatant before
and after the high-speed centrifugation will give an indication of how
much detergent was effect in the supernatant (membrane).
Reducing agents
Reducing agents are generally used in sample preparation to cleave
disulfide bond DTT, DTE and -ME are more commonly reducing
agents used. The aim of using these agents to protect proteins from
precipitating and aggregating due to oxidative crosslinking and to
reduce protein aggregation that may inhibit crystallization, Reducing
agents can interact with metals within protein sample and that is not
good for crystallization in this case. BME is particularly sensitive to
cobalt, copper and many phosphate buffers while DTT is sensitive to
nickel. But until now no specific article show different between the
DTT, DTE and -ME as well as in which crystallization step is better
to be use.
Isoelectric point (pI), pH & Temperature
The pH influence on protein solubility, different proteins
are soluble at different pH values, at high pH protein soluble
(deprotonate), low pH protein soluble (Protonated) and at isoelectric
point: protein aggregates. Acidic proteins has pI lower than 7, more
likely to crystallize around one pH unit above their pI, while basic
proteins more likely to crystallize lower than their pI about 1.53 pH
units [2]. Generally, protein aggregation or precipitation when the
solubility decrease at pH close to the isoelectric point (pI). It better to
move pH away from the pI to increase protein solubility and Super
saturation. At low salt concentration the electrostatic repulsions well
decrease when pH is different from pI, this well likely induce PEG to
effect depletion attraction. The depletion attraction due to polymer is
more effective when the protein net charge is lower.
To screen favor attraction between macromolecules and protein
charges it better to increasing ionic strength or gently moving pH
closer to pI. The most critical things is pI we need to make Sure the
protein having net charge close to zero and it soluble, and this depend
on some paper advice.
Actually, temperature is important parameter for crystallization,
it is better be screened. Temperature influences crystal growth and
nucleation by changing super saturation as well as solubility of the
sample. For instance, pH of Tris buffer is susceptible to temperature
changes. However, temperature can useful at low ionic strengths
by effects the amplified. Proteins very often show several crystal
026
polymorphs as result of different temperature. Lowering the growth
inducing temperature, this lessening the rate of protein synthesis and
usually more soluble protein is obtained. Because the temperature
can affects solubility is better to cool the Crystal solutions to prevent
protein degradation.
Optimum salt concentration
Different NaCl concentrations with different pH levels it was
use for crystallization. The effective of salt concentration in a
crystallization solution can be prognosticate before performing in the
experiment, because it is depended on the buffer pH and the pI of the
protein. It is important for successful crystallization to keep the salt
concentrations neither too low nor too high in the protein solution,
highly salt concentration may stabilize the protein buffer or decrease
the protein solubility lead to precipitation. Then crystallization
happens in the drop in which the components are concentrated
through water loss. If the original proteins as well as reservoir solution
do not contain enough salt, the concentration in the drop does not
reach the marginal concentration level.
Commonly, the increase of precipitant concentration in crystal
solutions is an effective technique adopted to lowering the solubility of
proteins. The salt it can stabilizes the water structure (Kosomtropic) or
disrupts the water structure (Chaotropic). In addition, the solubility
of the proteins drop at high concentrations of salt as well as solubility
of the proteins increases at low concentrations of salt. The optimal
salt concentration will slowly dehydrate our drop as well as increase
the concentration of our protein slowly.
The marginal ionic strength become high when the difference
between the pI of the proteins and the pH of the buffer is large, which
is agreement with the idea of the electrostatic screen effect of a salt [3].
Multi subunit proteins or virus it is large macromolecules,
highly salt concentrations are not effective to promote enough
interactions to induce crystallization while the Polyethylene glycol
(PEG) is a commonly used as precipitant in protein crystallization is
indispensable to induce crystallization.
Crystallization of iron protein under anaerobic condition may
facing precipitation problem, if the oxidation of the protein causing
the precipitation try to add sodium dithionite to preserve the reduced
state, also is better to change the ratio in the crystallization drop by
higher the amount of the precipitant.
Expression system
E coli is first choice for structural biologists to produce protein for
X-ray crystallography, but sometime unsuitable to express membrane
proteins in bacterial expression systems because the system cannot
provide the post-translational modifications, the folding machinery
and specific lipid environment. For eukaryotic membrane protein
better to express in Saccharomyces cerevisiae, Pichia pastoris, Sf9
insect cells [4] and human embryonic kidney.
Improve the expression level
In case of poor expression it need to be sure the protein is
not toxic for the cell. Choosing a smaller fragment of the target
protein can improve the expression levels and solubility, because
Citation: Abdalla M, Ali Eltayb W, Samad A, Elshareef SHM, Dafaalla TIM (2016) Important Factors Influencing Protein Crystallization. Glob J Biotechnol
Biomater Sci 2(1): 025-028.
Abdalla et al. (2016)
E. coli may not express well very large proteins > 70 kDa. Using
strains carrying mutations which remove the production of proteases
can occasionally enhance accumulation by diminution proteolytic
degradation, also reducing the growth temperature will produce
protein slow but at the same time slower proteolytic degradation.
Using special media containing trace metals (high density culture
media) it can increase protein yield 10 time over LB.
Co-expression of foldases proteins e.g. peptidyl prolyl cis/
trans isomerases (PPIs), disulfide oxidoreductase (DsbA), disulfide
isomerase (DsbC) and protein disulfide isomerase (PDI) with the
target protein may lead to highly levels of protein solubility. The other
advantages for this enzymes assist the formation of disulfide bonds
which does not happen in the reducing environment an also reduced
proteolysis
Commonly challenging and important task, protein aggregation
and precipitation is often happen when increasing a protein
concentration in the buffer to the required level.
Size exclusion chromatography
Is useful for further purification but some time is better to use
new column for purification. The shape of the chromatogram and
retention time provides.
High homogeneity and purity of the sample are crucial for the
crystallization to be successful, the presence of different aggregates or
oligomeric forms in the protein solution it well effect, we need to use
Dynamic light scattering (DLS) to check that or small-angle X-ray
scattering(SAXS).
Folding and stability
Natively unfolded proteins are some proteins unfolded in any
physiological conditions, and this kind may better to co crystallize
with other protein. UN folding, while highly beta strand are more
likely to form amyloid-like aggregates, but both of them can be
crystallize PDB (1aos--1by3). There is no direct correlation between
percentage of helix secondary structure as well as beta strand and
stability. However, proteins with high alpha-helical protein habitually
are less problematic upon. There are more hydrophobic contacts
in Beta sheets, misfolding or mutation might result in exposure of
hydrophobic residues. In this case, proteins aggregate to inhibit
hydrophobic exposure as well as avoid entropic penalty. Beta sheets
are stabilized through hydrogen bonding as well as hydrophobic
contacts, while Alpha helices are predominantly stabilized through
hydrogen bonding. Thus sheet is a little inferior in terms of stability.
Addition of co-factors or prostethic groups (such as a vitamin,
lipid, or inorganic like a metal ion) which are essential for protein
stability or for proper folding, but must be not fluctuation the pH
in the medium during growth, this leads to the accumulation of the
target protein osmoprotectants in the cell, which stabilize the protein
structure.
Biophysical methods like CD spectroscopy and other differential
scanning fluorometry (DSF) can be used for characterizing the
stability of the protein in different buffers, pH and in the presence
of different ligands to make sure that the protein is correctly folded.
There is several fold recognition websites (FOLDpro, RF-Fold,
SSHMM, THREADER, BLASTLINK, SSEARCH, PSI-BLAST and
HMMER) used to predict the protein fold.
Mutation
When the native protein fails to crystallize against ~ 300500
crystallization conditions it was propose genetically modified
proteins whether by truncations or point mutations might better for
crystallization.
Mutants of Tyr Phe and ThrVal more stable than the wild-type
protein, but have little effect on the conformation of the protein [5].
However, it has been reported that point mutations spectacular affect
the amount of aggregate formation in number of protein systems;
these include the interferon-, P22 tailspike protein, colicin A, singlechain antibodies, immunoglobulin domains, and interleukin-1 [6].
However, it is still not clear what mutations are likely to be helpful in
crystallization (Figure 1).
Cysteine and histidine predominantly react with reagents
prepared for the other and for other amino acid side chains as well,
that why it is problematic during crystallization. Some domain and
cloning are affecting the solubility this checking before expend time
on buffer optimization.
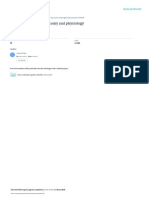
Figure 1: superdex 75 A. New superdex 75 B. Old superdex 75 as we can see new superdex 75 separate the protein to three pick, one is aggregation protein, two
is dimer protein, three is monomer, while the old superdex 75 cannot separate them and in this case is difficult to be crystal because the crystal condition for dimer
is not same to the monomer condition rather then that aggregation it can be crystal.
027
Citation: Abdalla M, Ali Eltayb W, Samad A, Elshareef SHM, Dafaalla TIM (2016) Important Factors Influencing Protein Crystallization. Glob J Biotechnol
Biomater Sci 2(1): 025-028.
Abdalla et al. (2016)
Nucleation
Sometime one protein can create different crystal, but that not
mean all of them they are stable, it can just be kinetically favored.
There is two kind of nucleation: heterogeneous is the most common,
occurs by interface of different composition and homogeneous as high
purity crystal. The nucleation rates it can be increased by increasing
the super saturation, solubility, viscosity and decreases temperature
and solidliquid interfacial tension.
Ethanol
In ethanol, the assist of peptide group burial to protein stability
would be increase and the contribution of non-polar group burial
would be decrease. Because -helices are expected to be stable
structures in ethanol, ethanol used to increase -helix formation in
proteins and peptides [7], also ethanol and methanol can lowers the
thermal denaturation temperature.
Glycerol
Often use in crystallization to protect the proteins as cryosolvent
because antifreeze properties and to enhance their solubility by
stabilizing the conformation, sugar protein specially affected by
glycerol because of ribonuleotides bonds. But some time glycerol
work as an antinucleation agent in crystallization.
TRIS & HEPES buffers
The differences in the influence of the buffers could be attributed
neither to disparity in the amounts of de-protonated buffer ions
nor to disparity in ionic strength. Tris and HEPES being positively
charged at pH 7.0. Protein activity increased in different buffer by
following order: citrate < Tris potassium phosphate <HEPES. While
oligonucleotide initiator decreased in the following order: cacodylate
> MES > HEPES > TRIS > phosphate [8]. Tris as well as HEPES
prevent the auto-oxidation. HEPES accelerated the degeneration rate
of the oxidant and peroxynitrite. The exist of as much as 100 mM
NaCl enhance the reversibility and stability of unfolding transitions
in Hepes buffer. The crystal structure obtain from HEPES buffer was
more similar to the active conformation.
Proteins in L-arginine and Tris buffer at pH 7.4 stay stable against
aggregation for longer periods of time. Tris is Goods buffers bind to
the DNA not only by electro static interactions but also by hydrogen
bonds primarily to the pyrimidine or purine rings [9]. Tris decrease
the metal ioncatalyzed oxidation of Cys, consequently, Cys being
available for interaction with the other reagent.
Conclusion
In this review presents important factors that affect protein
crystallization and ways to improve the results, but more needs to be
done to better understand. Some proteins have ability to make crystal
within initial concentration less than 1 mg per ml. Right working
sequence and buffer well make all the difference.
Acknowledgements
This work was supported by Ministry of Science and Technology
of China and the CAS-TWAS Presidents PhD Fellowship. All authors
declare no conflict of interests.
References
1. Golovanov AP, Hautbergue GM, Wilson SA, Lian LY (2004) A simple method
for improving protein solubility and long-term stability. J Am Chem Soc 126:
8933-8939.
2. Kirkwood J, Hargreaves D, OKeefe S, Wilson J (2015) Analysis of
crystallization data in the Protein Data Bank. Acta Crystallogr F Struct Biol
Commun 71: 1228-1234.
3. Yamanaka M, Inaka K, Furubayashi N, Matsushima M, Takahashi S, et
al. (2011) Optimization of salt concentration in PEG-based crystallization
solutions. J Synchrotron Radiat 18: 84-87.
4. Jasti J, Furukawa H, Gonzales EB, Gouaux E (2007) Structure of acidsensing ion channel 1 at 1.9 A resolution and low pH. Nature 449: 316-323.
5. Pace CN, Trevino S, Prabhakaran E, Scholtz JM (2004) Protein structure,
stability and solubility in water and other solvents. Philos Trans R Soc Lond B
Biol Sci 359: 1225-1234.
6. Izard J, Parker MW, Chartier M, Duche D, Baty D (1994) A single amino
acid substitution can restore the solubility of aggregated colicin A mutants in
Escherichia coli. Protein engineering 7: 1495-1500.
7. Luo P, Baldwin RL (1997) Mechanism of helix induction by trifluoroethanol:
a framework for extrapolating the helix-forming properties of peptides from
trifluoroethanol/water mixtures back to water. Biochemistry 36: 8413-8421.
8. Arzumanov AA, Victorova LS, Jasko MV, Yesipov DS, Krayevsky AA
(2000) Terminal deoxynucleotidyl transferase catalyzes the reaction of DNA
phosphorylation. Nucleic acids research 28: 1276-1281.
9. Righetti PG, Gelfi C, DAcunto MR (2002) Recent progress in DNA analysis
by capillary electrophoresis. Electrophoresis 23: 1361-1374.
Copyright: 2016 Abdalla M, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits
unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
028
Citation: Abdalla M, Ali Eltayb W, Samad A, Elshareef SHM, Dafaalla TIM (2016) Important Factors Influencing Protein Crystallization. Glob J Biotechnol
Biomater Sci 2(1): 025-028.
You might also like
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (119)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (265)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (587)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2219)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- Lab ValuesDocument3 pagesLab Valuessurviving nursing schoolNo ratings yet
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (890)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- Blood Pressure ChartDocument9 pagesBlood Pressure ChartVer BautistaNo ratings yet
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- The Progressive Muscle RelaxationDocument4 pagesThe Progressive Muscle Relaxationmaier_gabriela943980% (5)
- 01 - Newborn Physical ExamDocument2 pages01 - Newborn Physical Examgerald_valeriano0% (1)
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- How to Dilute Blood and Count White Blood CellsDocument2 pagesHow to Dilute Blood and Count White Blood CellsAlfred ChowNo ratings yet
- Pharmacology Toxicology - 4th EdDocument41 pagesPharmacology Toxicology - 4th EdYorlenie Juárez100% (1)
- Trauma y Memoria Procedural 2015 Paper Peter LevineDocument18 pagesTrauma y Memoria Procedural 2015 Paper Peter LevineMaría Regina Castro CataldiNo ratings yet
- Brain Organization and Memory - Cells, Systems, and Circuits PDFDocument429 pagesBrain Organization and Memory - Cells, Systems, and Circuits PDFMarvin Jay FarralesNo ratings yet
- NSTEMI Case PresentationDocument24 pagesNSTEMI Case PresentationMHIEMHOINo ratings yet
- Biological Effects of Microwaves and Radio Frequency RadiationDocument25 pagesBiological Effects of Microwaves and Radio Frequency RadiationMichael Janitch100% (2)
- P.E 2nd Quarter ExamDocument2 pagesP.E 2nd Quarter ExamCherry Vhim Flores Lanurias100% (3)
- Southern Blotting TechniqueDocument15 pagesSouthern Blotting TechniqueSouravS.PandaNo ratings yet
- A Survey of Solid Waste Management in Chennai (A Case Study of Around Koyambedu Market and Madhavaram Poultry Farms)Document4 pagesA Survey of Solid Waste Management in Chennai (A Case Study of Around Koyambedu Market and Madhavaram Poultry Farms)Peertechz Publications Inc.100% (1)
- Should National Environmental Policy Focus On Developing More Oil Resources or Developing Renewable Energy Sources?Document3 pagesShould National Environmental Policy Focus On Developing More Oil Resources or Developing Renewable Energy Sources?Peertechz Publications Inc.No ratings yet
- Comparative Analysis of Collapse Loads of Slabs Supported On Orthogonal Sided Frames With Beam/column Joint FixedDocument7 pagesComparative Analysis of Collapse Loads of Slabs Supported On Orthogonal Sided Frames With Beam/column Joint FixedPeertechz Publications Inc.No ratings yet
- Study On Improvement of Manpower Performance & Safety in Construction FirmsDocument5 pagesStudy On Improvement of Manpower Performance & Safety in Construction FirmsPeertechz Publications Inc.No ratings yet
- User Response - Based Sustainable Solutions To Traffic Congestion Problem Using Public Transport: The Case of Uttara, DhakaDocument7 pagesUser Response - Based Sustainable Solutions To Traffic Congestion Problem Using Public Transport: The Case of Uttara, DhakaPeertechz Publications Inc.No ratings yet
- Diagnosing HPV-Related Oropharyngeal Cancers: The Need To Speak A Common LanguageDocument4 pagesDiagnosing HPV-Related Oropharyngeal Cancers: The Need To Speak A Common LanguagePeertechz Publications Inc.No ratings yet
- Modulation of Immune Response in Edible Fish Against Aeromonas HydrophilaDocument5 pagesModulation of Immune Response in Edible Fish Against Aeromonas HydrophilaPeertechz Publications Inc.No ratings yet
- COMAMMOX - A New Pathway in The Nitrogen Cycle in Wastewater Treatment PlantsDocument3 pagesCOMAMMOX - A New Pathway in The Nitrogen Cycle in Wastewater Treatment PlantsPeertechz Publications Inc.No ratings yet
- The Role of Decavanadate in Anti - Tumour ActivityDocument3 pagesThe Role of Decavanadate in Anti - Tumour ActivityPeertechz Publications Inc.No ratings yet
- A Perceptive Journey Through PostmodernismDocument4 pagesA Perceptive Journey Through PostmodernismPeertechz Publications Inc.No ratings yet
- Is It Time For The Introduction of Colostomy Free Survival in Rectal Cancer PatientsDocument4 pagesIs It Time For The Introduction of Colostomy Free Survival in Rectal Cancer PatientsPeertechz Publications Inc.No ratings yet
- Primary Spindle Cell Rhabdomyosarcoma of Prostate: A Case Report With Review of LiteratureDocument3 pagesPrimary Spindle Cell Rhabdomyosarcoma of Prostate: A Case Report With Review of LiteraturePeertechz Publications Inc.No ratings yet
- Abrikossoff's Tumour Mimicking A Neoplastic Tumour of The Breast: A Case ReportDocument3 pagesAbrikossoff's Tumour Mimicking A Neoplastic Tumour of The Breast: A Case ReportPeertechz Publications Inc.No ratings yet
- Role of Tumor Heterogeneity in Drug ResistanceDocument2 pagesRole of Tumor Heterogeneity in Drug ResistancePeertechz Publications Inc.No ratings yet
- Angiosarcoma of The Scalp and Face: A Hard To Treat TumorDocument4 pagesAngiosarcoma of The Scalp and Face: A Hard To Treat TumorPeertechz Publications Inc.No ratings yet
- MEK Inhibitors in Combination With Immune Checkpoint Inhibition: Should We Be Chasing Colorectal Cancer or The KRAS Mutant CancerDocument2 pagesMEK Inhibitors in Combination With Immune Checkpoint Inhibition: Should We Be Chasing Colorectal Cancer or The KRAS Mutant CancerPeertechz Publications Inc.No ratings yet
- Echinocandins and Continuous Renal Replacement Therapies: The Role of AdsorptionDocument3 pagesEchinocandins and Continuous Renal Replacement Therapies: The Role of AdsorptionPeertechz Publications Inc.No ratings yet
- The Promise of Disease Management in GreeceDocument8 pagesThe Promise of Disease Management in GreecePeertechz Publications Inc.No ratings yet
- PLGF, sFlt-1 and sFlt-1/PlGF Ratio: Promising Markers For Predicting Pre-EclampsiaDocument6 pagesPLGF, sFlt-1 and sFlt-1/PlGF Ratio: Promising Markers For Predicting Pre-EclampsiaPeertechz Publications Inc.No ratings yet
- Body Mass Index Impact and Predictability On Preeclamptic ToxemiaDocument6 pagesBody Mass Index Impact and Predictability On Preeclamptic ToxemiaPeertechz Publications Inc.No ratings yet
- Video Endoscopic Inguinal Lymphadenectomy: Refi Ning Surgical Technique After Ten Years ExperienceDocument4 pagesVideo Endoscopic Inguinal Lymphadenectomy: Refi Ning Surgical Technique After Ten Years ExperiencePeertechz Publications Inc.No ratings yet
- Steroid Withdrawal Protocols in Renal TransplantationDocument8 pagesSteroid Withdrawal Protocols in Renal TransplantationPeertechz Publications Inc.No ratings yet
- An Overview: Laboratory Safety and Work Practices in Infectious Disease ResearchDocument6 pagesAn Overview: Laboratory Safety and Work Practices in Infectious Disease ResearchPeertechz Publications Inc.No ratings yet
- Bacterial Vaginosis: Risk of Adverse Pregnancy OutcomeDocument3 pagesBacterial Vaginosis: Risk of Adverse Pregnancy OutcomePeertechz Publications Inc.No ratings yet
- Evaluating Mechanical Ventilation in Patients With Acute Respiratory Distress SyndromeDocument5 pagesEvaluating Mechanical Ventilation in Patients With Acute Respiratory Distress SyndromePeertechz Publications Inc.No ratings yet
- The Outcomes of Postoperative Total Hip Arthroplasty Following Western Ontario McMaster Universities Osteoarthritis Index (WOMAC) : A Prospective StudyDocument6 pagesThe Outcomes of Postoperative Total Hip Arthroplasty Following Western Ontario McMaster Universities Osteoarthritis Index (WOMAC) : A Prospective StudyPeertechz Publications Inc.No ratings yet
- The Outcomes of Postoperative Total Hip Arthroplasty Following Western Ontario McMaster Universities Osteoarthritis Index (WOMAC) : A Prospective StudyDocument6 pagesThe Outcomes of Postoperative Total Hip Arthroplasty Following Western Ontario McMaster Universities Osteoarthritis Index (WOMAC) : A Prospective StudyPeertechz Publications Inc.No ratings yet
- An Overview: Laboratory Safety and Work Practices in Infectious Disease ResearchDocument6 pagesAn Overview: Laboratory Safety and Work Practices in Infectious Disease ResearchPeertechz Publications Inc.No ratings yet
- Exercise-Induced Time-Dependent Changes in Plasma BDNF Levels in People With SchizophreniaDocument6 pagesExercise-Induced Time-Dependent Changes in Plasma BDNF Levels in People With SchizophreniaPeertechz Publications Inc.No ratings yet
- Exercise-Induced Time-Dependent Changes in Plasma BDNF Levels in People With SchizophreniaDocument6 pagesExercise-Induced Time-Dependent Changes in Plasma BDNF Levels in People With SchizophreniaPeertechz Publications Inc.No ratings yet
- Fundamentals of Respiratory Sounds and AnalysisDocument68 pagesFundamentals of Respiratory Sounds and AnalysisVaidotas NorkūnasNo ratings yet
- Blood Supply of BrainDocument49 pagesBlood Supply of BrainDarling Sevenfoldism SynysterNo ratings yet
- Cellular ComponentsDocument2 pagesCellular ComponentsWangSheng TanNo ratings yet
- Physical BackgroundDocument24 pagesPhysical BackgroundmarkNo ratings yet
- Subject Enrichment Activity For Term 2Document5 pagesSubject Enrichment Activity For Term 2AADITRYA JAINNo ratings yet
- Unit I Clothing Science Two Marks With Answer and Question BankDocument3 pagesUnit I Clothing Science Two Marks With Answer and Question BankSivakumar KNo ratings yet
- Biology Savemyexam NotesDocument4 pagesBiology Savemyexam NotesSayeef MahdiNo ratings yet
- Synevo Results 5Document3 pagesSynevo Results 5Kenan BagirNo ratings yet
- HydrochlorothiazideDocument3 pagesHydrochlorothiazideapi-3797941No ratings yet
- Microsyllabus BSC 301Document9 pagesMicrosyllabus BSC 301Anwita JhaNo ratings yet
- Cormie2011 PDFDocument22 pagesCormie2011 PDFGust AvoNo ratings yet
- Acute Renal Failure in The Intensive Care Unit: Steven D. Weisbord, M.D., M.Sc. and Paul M. Palevsky, M.DDocument12 pagesAcute Renal Failure in The Intensive Care Unit: Steven D. Weisbord, M.D., M.Sc. and Paul M. Palevsky, M.Dkerm6991No ratings yet
- The Ghost Map NotesDocument9 pagesThe Ghost Map NotesLuke CybulskiNo ratings yet
- Auditory Pathways: Anatomy and Physiology: Handbook of Clinical Neurology March 2015Document24 pagesAuditory Pathways: Anatomy and Physiology: Handbook of Clinical Neurology March 2015florensiaNo ratings yet
- من جامعة الأردن PDFDocument266 pagesمن جامعة الأردن PDFyousefNo ratings yet
- Gastrointestinal Diseases: Psycho-Social Aspects: BackgroundDocument7 pagesGastrointestinal Diseases: Psycho-Social Aspects: BackgroundPedro LosadaNo ratings yet
- Osmosis Lab DiscussionDocument2 pagesOsmosis Lab Discussionsoha zubairNo ratings yet
- ACOSTA, Joyce Ara, SBAR 3 (In Good Condition)Document2 pagesACOSTA, Joyce Ara, SBAR 3 (In Good Condition)Joyce Ara Tumbaga AcostaNo ratings yet