Professional Documents
Culture Documents
Ultra Performance Liquid Chromatographic Method For Quantification of Clofarabine Related Substances
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Ultra Performance Liquid Chromatographic Method For Quantification of Clofarabine Related Substances
Copyright:
Available Formats
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Ultra-Performance Liquid Chromatographic Method for
Quantification of Clofarabine Related Substances in an
Injection Formulation
Dhananjay Prasad Dwivedi* B Ravi Kumar,Konda Sidda Reddy, Prof.J. Sreeramlu
*Department of Chemistry Rayalaseema
University Kurnool- Andhra Pradesh, India
Abstract:-A new and stability-indicatingreversed- 4-fluoro-2- (hydroxymethyl) oxolan-3-ol empirical
phase ultra performance liquid formula is CioHi IC1 FN503 and molecular weight is
chromatographymethod was developed and 303.67 g/mol (Figure 1). Clofarabine developed by
validated for simultaneous determination of Genzyme Corporation, USA, listed on January 11, 2005
clofarabine impurities in Injectionformulation. The by the U.S. FDA expedited procedures, as over the past
chromatographic separation was carried out on an decade the first approved specifically for the treatment
ACQUITY UPLC CSH C18 (50mm x 2.1mm, of childhood leukemia drug, mainly used in the
1.7m)using a mobile phase consisting of treatment of children with refractory or recurrent acute
Ammonium formate buffer with pH 3.0 lymphoblastic leukemia (ALL) in children previously at
andacetonitrile at a flow rate of 0.28 ml/min and an least has received two therapy invalid. Clofarabine is
injection volume of 1L. The method was validated not listed in any pharmacopoeia, however few
for precision, accuracy, specificity, linearity, chromatographic methods are reported on its
sensitivity and robustness. The UV detectio n was determination. Recently, LC-MS/MS method was used
performed at 264 nm.The proposed method can be for determination of clofarabine in human urine and
applied for quality control, release and stability plasma. Of late, HPLC method is reported for the
analysis of clofarabine and its impurities in determination of concentration of clofarabine in rat
Injection formulation. plasma [10]. LC/MS method is reported for analysis
and identification of chlorinated impurity of
clofarabine. Ultra performance convergence
Keywords:- Clofarabine, Stability-indicating, UPLC, chromatography method was used for identification of
Validation. two related substances in clofarabine [11]. Over period,
several stabilityindicating reverse phase high
I. IN T RO D UC T IO N: performance liquid chromatography (RP-HPLC)
methods were developed for determination of
Clofarabine, an antimetabolite purine nucleoside used concentration of clofarabine [14-15].
in the treatment of lymphoblastic leukemia. The
clofarabine is intracellularly converted to 5' - II. CH EM ICA L S T RU C TU RE O F
monophosphate by deoxycytidine kinase and 5'- CL O FA RAB I N E A ND I T S IM PUR I TI E S :
triphosphate by mono- and di-phosphokinases [101.
This metabolite halts DNA synthesis by inhibiting
ribonucleotide reductase leading to depletion of
intracellular deoxynucleotide triphosphate pool.
However, self - potentiation of clofarabine triphosphate
incorporation into DNA increases the extent of
inhibition of DNA synthesis [Ill Preclinical models has
been demonstrated that clofarabine 5'- triphosphate
inhibits DNA repair by incorporation into the DNA
chain during the repair process [12-13]. It disrupts the
mitochondrial membrane integrity and induces
apoptosis by releasing cytochrome C and apoptosis -
inducing factor [11. The IUPAC name of clofarabine a) Clofarabine
is (2R, 3R, 4S, 5R)-5-(6-amino- 2-chloropurin-9- y1)-
IJISRT17JL171 www.ijisrt.com 334
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
B. Selection of detection wavelength
The sensitivity of a method that uses a UV detector depends on
the proper selection of wavelength. An ideal wavelength is that
which is maximally absorbed and provides an acceptable
response for the drug, which should not interfere with other
peaks.
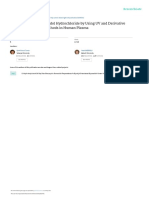
UV spectra of the drug and its impurities were recorded by
b) Impurity-A: scanning between 200 and 400 nm.The spectra of drug and its
impurities were overlaid and the wavelength 264 nm was
selected where the active analyte as well as impurities have
sufficient response for detection and quantification. The
ultraviolet scans of Clofarabine and the three potential
impurities are depicted.
C. Instrument and Chromatographic condition:
The Integrated Acquity UPLC system used for the study was
purchased from Waters Corporation, Milford, USA and
equipped with Waters photodiode array detector (PDA). Data
collection and analysis was performed using Empower
software 2pro (Waters Corporation). The balance used for
weighing the reference standards and samples was purchased
from Metter Toledo. Separation was achieved on a Waters
c) Impurity-B:
acquity CSH C18 column with dimensions 50 mm x 2.1 mm
I.D and a particle size of 1.7 m. A simple mobile phase
consisting of Ammonium formate buffer (0.01 M, pH 3.0) and
Buffer: acetonitrile (Mobile phase B) was pumped into the
UPLC chromatograph using a gradient program with varying
compositions at a flow rate of 0.28 mL/ min, with a column
temperature of 50C throughout the run. Sample volume of
1pL was injected into the chromatograph and detected at 264
nm.
The final conditions summarized in the Table.No:1 and 2.
Preparation of standard and sample solution
d) Impurity-C: A mixture of water and methanol in the ratio 20:80 was used as
a diluent for preparing the solutions of standard and samples
(diluent).
Standard stock solution
III. MATERIALS AND METHODS
A standard stock solution was prepared by dissolving 25 mg of
Clofarabine working standard in 100 mL of the diluent.
A. Chemicals And Drugs
Preparation of standard solution for impurities
Ammonium formate (AR Grade) was used to prepare
determination (0.001 mg per mL)
the buffer and obtained from Sigma -Aldrich. Formic
acid (AR Grade) used to adjust the pH, was obtained
The standard solution for the determination of impurities was
from Merck Specialties, India. The active
prepared by diluting 5 mL of the standard stock into 50 mL with
pharmaceutical ingredient clofarabine and impurities
the diluent and further diluted 2 mL of the solution into 50 mL
were obtained from internally.
with the diluent to obtain a concentration of 0.001 mg per mL.
IJISRT17JL171 www.ijisrt.com 335
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Preparation of sample and placebo solution for impurities Preparation of spiked sample solution for impurities
determination. determination
Sample solution was prepared by diluting 2 mL of the pooled Stock solution of all impurities was prepared by dissolving an
Clofarabine injection to 10 mL using the diluents to obtain a appropriate quantity of each impurity in the diluent to obtain a
concentration of 0.5 mg per mL. concentration of 1 mg per mL. An appropriate volume of
impurity stock solution was diluted with sample solution to get
Placebo equivalent to 5 mg of the sample was taken and diluted a final concentration of 0.2% for each impurity.
to 10 mL with the diluents and mixed.
210.0210.0
263.0 Clofarabine
264.0 Impurity B
264.0
Impurity A
263.0
Impurity C
336.0 370.0 389.0
328.0 375.0
200.00 220.00 240.00 260.00 280.00 300.00 320.00 340.00 360.00 380.00 400.00
nm
Figure 2.
D. Method development and optimization
and their acceptable limits was collected to define sample
Clofarabine has ionizable amino groups, and hence, the concentration and the range of the method. The maximum daily
retention time of the drug is highly dependent on the pH of the dose of Clofarabine is 50mg/day. Based on the daily dose, the
mobile phase. In the present study, the pH of the mobile phase qualification threshold did not exceed 0.5%, and the
was maintained acidic (pH = 3.0) by the addition of formic acid identification threshold was 0.2%. Method development was
solution. This yields narrow and symmetrical peak.Before targeted to cover a range of 50-150% of identification threshold
initiating the development activity, information on impurities for impurities. A systematic approach was adopted for the
method development.
Fig . 3 . Chromatogram obtained from Trial-1.
IJISRT17JL171 www.ijisrt.com 336
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Another attempt was made by changing the gradient program parameters were unchanged. All impurities were appropriately
and keeping other parameters unchanged. The patterns obtained separated from each other. The resolution between impurity B
from both trials did not differ significantly. The pattern obtained and the Clofarabine is further improved by modifying the
for trial 2 is depicted in Figure 4. gradient program. The optimized chromatographic conditions
are listed in Table:1 and 2. A specimen chromatogram obtained
The next trial was made by changing the column to Waters CSH from the final method parameters is illustrated in Figure 5.
C18, 50 x 2.1 mm, 1.7 m. The remaining chromatographic
Fig . 4 . Chromatogram obtained from Trial-2.
Fig . 5 . Specimen Chromatogram obtained from final method:
IJISRT17JL171 www.ijisrt.com 337
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Table No. : 1. Optimized Chromatographic Conditions.
Chromatograph WatersAcquity UPLC system
Mobile Phase A: pH3.0, 10mM Ammonium formate Buffer
Mobile phase Mobile Phase B: Buffer : Acetonitrile (20:80v/v)
(80:20%v/v)
Acquity UPLC CSH C18, (50mmx2.1mm) 1.7m
Column
size)
Detector PDA
Flow rate 0.28 ml/min
Wavelength detection 264 nm
Injection volume 1L
Column Oven Temperature 50C
Run time 16 min
Diluent Methanol: Water (80:20 v/v)
Table No. : 2. Gradient Programmed.
Time(mins) Flow(ml/min) %A %B
Initial 0.28 95.00 5.00
5.00 0.28 95.00 5.00
7.00 0.28 75.00 25.0
9.00 0.28 60.00 40.0
12.00 0.28 15.00 85.0
14.00 0.28 95.00 5.00
16.00 0.28 95.00 5.00
IJISRT17JL171 www.ijisrt.com 338
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
IV. METHOD VALIDATION
Table No. : 3. System suitability data and Precision data.
A. Results and Discussion
Peak Area count
The optimized method was fully validated for determination of
Preparatio
impurities as per ICH guidelines, (Q2A (R1) validation of Impurit
analytical procedures. The method was validated for specificity, n Clofarabi Impurit Impurit
precision, accuracy, linearity,Limit of Detection (LOD), Limit y-C
ne y-A y-B
of Quantification (LOQ) and Robustness.
Peak
System Suitability.
Tailing 1.1 1.0 1.2 1.1
System suitability parameters were measured to verify the
system performance for the intended analysis. Hence, system
Theoretical
precision was determined on six replicate injections of standard
preparations and the relative standard deviation (% RSD) was plates 7271 10984 6542 12549
evaluated. In addition to the % RSD, USP tailing, and USP plate
count were also evaluated and found to be satisfactory, as per
current USP requirements for a chromatographic peak (USP %RSD 1.9 0.8 1.4 0.8
General Chapter<621>). The system suitability results obtained
are presented in Table 3. Accuracy
Precision Accuracy of the analytical procedure expresses the degree of
closeness of the obtained results with true values. Samples for
Precision of the test method was evaluated by injecting six accuracy were prepared in triplicate by spiking impurities at
individual samples spiked with all three impurities into the different levels.. The covered levels for impurities were LOQ,
chromatograph. The % RSD values from the six individual test 50% , 100% and 150% of the Identification Threshold (0.2%)
preparations were found to be below 2% for all three impurities. with respect to the sample concentration (0.5mg/ mL) From the
The ruggedness (intermediate precision) of the method was response of the analyte peaks, the amounts recovered (in %) and
determined using another system and column for the analysis in % RSD were reported. Accuracy results are summarized in
a different day. The precision data are listed in Table 3. The Table 4.
data indicate that a low % RSD, concluding that the method is
precise for impurities determination. The data indicate that the % recovery of impurities lies in the
range of 89.5-106.4 with an RSD of between 0.8 and 3.4%. This
indicates that the method is accurate and precise.
Table No. : 4. Recovery of Impurities data
Recovery Level (%)
Impurity Name
LOQ %RSD 50 %RSD 100 %RSD 150 %RSD
104.7 95.5 97.7 95.3
Impurity-A 99.7 3.4 99.7 2.3 93.8 3.1 92.2 1.7
106.4 96.2 99.7 93.3
94.0 98.7 103.2 95.2
Impurity-B 89.5 2.5 95.3 2.2 102.2 0.9 96.5 2.8
92.3 99.4 104.0 100.5
104.2 93.3 100.2 95.5
Impurity-C 98.5 3.0 92.5 0.8 101.8 1.0 96.7 2.3
103.2 94.0 102.1 92.4
IJISRT17JL171 www.ijisrt.com 339
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Linearity
Limit of Detection and Limit of Quantization.
The linearity of the analytical method was tested to check its
The LOD and LOQ values were established by Signal to Noise ability to elicit test results that are directly proportional to the
ratio. Results are shown in Table no.5 concentration of analyte in samples within a given range.
Hence, different concentrations of individual impurities and
Table No. : 5. Impurities LOD and LOQ Concentrations. standard working solution of Clofarabine were prepared and
injected into the UPLC, and the chromatograms were recorded.
Data based on signal to noise ratio (S/N) The linearity of detector response was determined by plotting a
Impurity graph of peak areas versus concentrations. Plots of linearity
LOQ experiments are illustrated in Figure 6. The correlation
name LOD Concentration
Concentration coefficients, >0.998 for impurities and 0.997 for Clofarabine,
(%w/w)
(%w/w) indicate that the method satisfies a linear relationship between
0.01143 the concentration and peak response. The linearity range is be-
Impurity A 0.0381 tween the limit of quantification (LOQ) and 150% of the sample
Impurity B 0.0524 0.01572 concentration for all impurities. The linearity data for all four
impurities and Clofarabine are listed in Table 6 to 9.
Impurity C 0.0421 0.01263
Clofarabine
20000
y = 10420x + 511.4
15000
Peak Area
R = 0.9979
10000
5000
0
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6
Concentration(g/mL)
Fi g.6 . C lo far ab i n e
Impurity A
40000
30000 y = 19730x + 1032.4
Peak Area
R = 0.999
20000
10000
0
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6
Concentration(g/mL)
Fi g. 7 I mp uri t y A .
IJISRT17JL171 www.ijisrt.com 340
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Impurity B
10000
y = 5994.5x + 15.788
8000
R = 0.9981
Peak Area
6000
4000
2000
0
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6
Concentration(g/mL)
Fi g. 8 . I mp uri t y B
Impurity C
14000
12000 y = 7524.5x + 565.39
10000 R = 0.9991
Peak Area
8000
6000
4000
2000
0
0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6
Concentration(g/mL)
Fi g. 9 . I mp uri t y C
power of the method. Specificity was determined by exposing
Specificity test solution to oxidation by hydrogen peroxide, acid hydrolysis,
base hydrolysis, heat and photolytic stress. A detailed procedure
The specificity of the RP-HPLC method was checked by
has been reported.
comparison of chromatogram obtained from standard, placebo
and sample. The Placebo did not should not show any
The stressed samples were then further diluted with the diluent
interference at retention of main analyte peak and at impurities.
and chromatographed as per the proposed method.
The peak purity of Clofarabine peak was evaluated using the
Forced degradation study.
PDA. The purity angle should be less than the purity threshold.
Forced degradation studies were conducted on samples and on The results are summarized in Table.No:6
the plain placebo to prove the specificity and stability-indicating
IJISRT17JL171 www.ijisrt.com 341
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Table No. : 6. Degradation Data.
Peak purity
Stress condition %degradation Purity flag
Purity Angle Purity Threshold
Treated with 5N HCL solution for
24hrs on bench top. 5.42 0.10 0.24 NO
Treated with 5N NaOH solution for
24hrs on bench top. 11.7 0.11 0.23 NO
Treated with 30% H2O2 solution for
24hrs on bench top. 0.40 0.10 0.24 NO
Treated with heat at 60C for 72hrs.
0.38 0.27 0.52 NO
Exposed to sunlight 1.2 million lux
hrs for 6days.
0.09 0.24 0.62 NO
Exposed to UV-Light 200
watts/square meter for 6days. 0.05 0.18 0.32 NO
Fi g. 1 0 . Acid Sample.
IJISRT17JL171 www.ijisrt.com 342
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Fi g. 1 1 . Base Sample.
Fi g. 1 2 .Thermal Sample.
Fi g. 1 3 . Sunlight Sample.
IJISRT17JL171 www.ijisrt.com 343
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
Robustness. results of the method. Result are shown in Table no.8 The
robustness of an analytical procedure is a measure of its
Reliability during normal usage. The robustness of a method is capacity to remain unaffected by small, but deliberate variations
evaluated by varying method parameters such as mobile phase in method parameters and provides an indication of its
composition, pH, etc., and determiningthe effect (if any) on the
Table No. : 7. Robustness conditions.
Parameters Change of conditions
Flowvariation High(+) 0.31
(ml/min)
Actual 0.28
Low(-) 0.25
Buffer pH High(+) 3.2
variation
Actual 3.0
Low(-) 2.8
Column oven temp. High(+) 55
variation (C)
Actual 50
Low(-) 45
Organic phase (%) High(+) 88
Mobile phase B Actual 80
Low(-) 72
Table No. : 8. Robustness data.
Relative Retention Time (RRT)
Impurity
Name Control pH(+) pH(-) Flow(+) Flow(-) Temp(+) Temp(-) Organic(+) Organic(-)
Impurity A 0.28 0.27 0.28 0.28 0.28 0.28 0.28 0.28 0.28
Impurity B 1.18 1.12 1.16 1.16 1.16 1.16 1.16 1.16 1.16
Impurity C 1.92 1.75 1.86 1.86 1.86 1.86 1.86 1.86 1.86
IJISRT17JL171 www.ijisrt.com 344
Volume 2, Issue 7, July 2017 International Journal of Innovative Science and Research Technology
ISSN No: - 2456 2165
[13]. L. J. De Gennero, E. Raetz, Acute lymphoblastic
V. CONCLUSIONS leukemia (Leukemia and Lymphoma Society, 2014, pp
1-52).
The rapid gradient reverse phase UPLC method, developed for
[14]. Brijesh D Patel, Usmangani K Chhalotiya, Dhruv B
the quantitative analysis of clofarabine related substances in
Patel Quantification of Newer Anti-Cancer Drug
pharmaceutical dosage forms, is precise, accurate, linear,
Clofarabine in their Bulkand Pharmaceutical Dosage
specific, and robust. Satisfactory results were obtained from
Form.J Chromatogr Sep Tech 2016, 7:3.
validation of the method. The method is stability-indicating and
[15]. B.D.Patel, U. K.Chhalotiya, D.B.Patel,
can be used for routine analysis of production samples, and
Quantification of Newer Anti-Cancer Drug Clofarabine
checking the stability of samples of Clofarabine Injection.
in their Bulk and Pharmaceutica Dosage Form. J
Chromatogr Sep Tech, 7(3), 2016, 328-832.
REFERENCES
[1]. Chemical and Physical properties of Clofarabine
(2004) Chem Spider.
[2]. DeGennero LJ, Raetz E (2014) Acute Lymphoblastic
Leukemia.
[3]. Fanali, S., Haddad, P., Poole, C., Schoenmakers, P., &
Lloyd, D.(2013). Liquid chromatography. Waltham,
MA: Elsevier.
[4]. Q2A (R1) validation of analytical procedures: Text and
methodology, international conference on
harmonization.2005.
[5]. Drug Bank. Retrieved from
http://www.drugbank.ca/drugs/DB00631.
[6]. Bakshi M, Singh S, Development of validated stability
indicating assay methods critical review. J Pharm
Biomed. 2002:28:1011-1040.
[7]. C.H.Pui, S. Jeha, P.Kirkpatrick, Clofarabine, Nature
Reviews Drug Discovery, 4, 2005, 369-370.
[8]. .L. Peng, T. Farkas, Analysis of basic compounds by
reversed-phase liquid chromatography electrospray mass
spectrometry in high-pH mobile phases, J Chromatogr
A., 1179, 2008,131144.
[9]. A. Zhenchuka, K. Lotfib, G. Juliussond, F. Albertionia,
Mechanisms of anti-cancer action and pharmacology of
clofarabine, Biochemical Pharmacology, 78(11),
2009,13511359.
[10]. M.A. Zhang-Qing, H.Zong-Yuan, L. Jia-Jie.
Determination of clofarabine in rat plasma and the
pharmacokinetic research, ActaMetallurgica Sinica,
16(8), 2011, 852-855.
[11]. N.R.Venkata NR, D. Jalandhar, G. Gnanadey, R.
Bandari, P. Manoi, Development of supercritical Fluid
(Carbon dioxide) based ultra-performance convergence
chromatographic stability indicating assay method for
determination of clofarabine in injection.Analytical
methods, 5, 2013, 7008-7013.
[12]. L. Huang, P. Lizak, C. Dvorak, J. Long-Boyle,
Simultaneous determination of Fludarabine and
Clofarabine in human plasma by LC-MS/MS, Journal of
Chromatography B Analyt Technol Biomed Life Sci.,
960, 2014, 194-199.
IJISRT17JL171 www.ijisrt.com 345
You might also like
- Analytical Methods for Major and Modified Nucleosides - HPLC, GC, MS, NMR, UV and FT-IRFrom EverandAnalytical Methods for Major and Modified Nucleosides - HPLC, GC, MS, NMR, UV and FT-IRNo ratings yet
- Analytical Methods: PaperDocument6 pagesAnalytical Methods: PaperRatnakaram Venkata NadhNo ratings yet
- Analytical Methods: PaperDocument6 pagesAnalytical Methods: PaperHeidi HughesNo ratings yet
- Admin, Journal Manager, 24199-118524-1-CEDocument5 pagesAdmin, Journal Manager, 24199-118524-1-CESachin BagewadiNo ratings yet
- 10.1515 - Revac 2022 0039Document12 pages10.1515 - Revac 2022 0039yordanosezerihun07No ratings yet
- Determination of Morphine in Human Urine by A Simple Reverse Phase High-Performance Liquid Chromatography Method With UV DetectionDocument5 pagesDetermination of Morphine in Human Urine by A Simple Reverse Phase High-Performance Liquid Chromatography Method With UV DetectionAdi SubagioNo ratings yet
- Validation of High-Performance Liquid Chromatographic-Mass Spectrometric Method For The Analysis of Lidocaine in Human PlasmaDocument4 pagesValidation of High-Performance Liquid Chromatographic-Mass Spectrometric Method For The Analysis of Lidocaine in Human PlasmaDiego RodriguezNo ratings yet
- Metodo Isocratico Por HPLC de Besifloxacino HCLDocument5 pagesMetodo Isocratico Por HPLC de Besifloxacino HCLJuan SNo ratings yet
- Development and Validation of RP-HPLC Method For Simultaneous Estimation of Ivermectin and Clorsulon in Ivercam InjectionDocument9 pagesDevelopment and Validation of RP-HPLC Method For Simultaneous Estimation of Ivermectin and Clorsulon in Ivercam InjectionFaelFernandesNo ratings yet
- Development and Validation of A HPTLC Method For Rivaroxaban in Human Plasma For A Pharmacokinetic StudyDocument6 pagesDevelopment and Validation of A HPTLC Method For Rivaroxaban in Human Plasma For A Pharmacokinetic Studypramod aloorNo ratings yet
- UV Derivative Spectrophotometric Analysis of SpironolactoneDocument6 pagesUV Derivative Spectrophotometric Analysis of SpironolactoneJay RanaNo ratings yet
- Chiral Separation and Modeling of Quinolones On Teicoplanin Macrocyclic Glycopeptide Antibiotics CSPDocument8 pagesChiral Separation and Modeling of Quinolones On Teicoplanin Macrocyclic Glycopeptide Antibiotics CSP5netNo ratings yet
- Determination of Spectinomycin Hydrochloride and Its Related Substances by HPLC-ELSD and HPLC-MSDocument5 pagesDetermination of Spectinomycin Hydrochloride and Its Related Substances by HPLC-ELSD and HPLC-MSMichael GaniNo ratings yet
- HPLC Method for Zinc Carnosine AnalysisDocument5 pagesHPLC Method for Zinc Carnosine AnalysisSouheila MniNo ratings yet
- 1 s2.0 S1110093112000312 MainDocument11 pages1 s2.0 S1110093112000312 Maindanielafarmacie_1617No ratings yet
- New UV Method for Chlorpheniramine Maleate AnalysisDocument6 pagesNew UV Method for Chlorpheniramine Maleate AnalysisArikus Wibowo MuhammadNo ratings yet
- Pengujian MorfinDocument7 pagesPengujian MorfinShofia CahyaNo ratings yet
- Automated Analysis For Free and Short-Chain Acylcarnitine in Plasma With A Centrifugal AnalyzerDocument6 pagesAutomated Analysis For Free and Short-Chain Acylcarnitine in Plasma With A Centrifugal AnalyzerHarry YucraNo ratings yet
- Articulo 1 SalchichasDocument5 pagesArticulo 1 SalchichasLALILA ORTIZ DIAZNo ratings yet
- Bai Bao EC-1998Document5 pagesBai Bao EC-1998Bình GiangNo ratings yet
- TLC Chromatographic-Densitometric Assay of Ibuprofen and Its ImpuritiesDocument5 pagesTLC Chromatographic-Densitometric Assay of Ibuprofen and Its ImpuritiesCarla MekdadNo ratings yet
- 3845-Article Text-10897-2-10-20200114Document5 pages3845-Article Text-10897-2-10-20200114nhan phamNo ratings yet
- Chapter 2Document5 pagesChapter 2UpadhayayAnkurNo ratings yet
- Art 08 PDFDocument7 pagesArt 08 PDFfarooq shikhNo ratings yet
- Validated HPLC Method for Quetiapine ImpuritiesDocument9 pagesValidated HPLC Method for Quetiapine ImpuritiesVinaya SnehalathaNo ratings yet
- Rapid RP-HPLC Method For The Quantification of Glabridin in Crude Drug and in Polyherbal FormulationDocument6 pagesRapid RP-HPLC Method For The Quantification of Glabridin in Crude Drug and in Polyherbal Formulationahmetgezer34No ratings yet
- Optimization of micellar LC for opium alkaloid analysisDocument9 pagesOptimization of micellar LC for opium alkaloid analysisartemNo ratings yet
- Multianalyte Serum Analysis Using Mid-Infrared SpectrosDocument9 pagesMultianalyte Serum Analysis Using Mid-Infrared SpectrosGutoGonçalvesNo ratings yet
- Ijppr 2021Document16 pagesIjppr 2021Smita NayakNo ratings yet
- BORNEOLDocument6 pagesBORNEOLKhánh ToànNo ratings yet
- A Hplc-Uv Method For Deteermination of Three Pesticides in WaterDocument8 pagesA Hplc-Uv Method For Deteermination of Three Pesticides in WaterHasna NoerNo ratings yet
- Experiment No. 6_ ThinLayerChromatography.docxDocument5 pagesExperiment No. 6_ ThinLayerChromatography.docxKarla Joy P. SucgangNo ratings yet
- Application of Liquid Chromatography Nuclear Magnetic Resonance Spectroscopy To The Identification of Natural ProductsDocument9 pagesApplication of Liquid Chromatography Nuclear Magnetic Resonance Spectroscopy To The Identification of Natural ProductsJuan Camilo ZuluagaNo ratings yet
- Deferiprone A Review of Analytical MethodsDocument7 pagesDeferiprone A Review of Analytical Methodssunilvarma3112No ratings yet
- Jaoac 0311Document11 pagesJaoac 0311adolfo olmosNo ratings yet
- J Jtusci 2014 06 001Document7 pagesJ Jtusci 2014 06 001Mohamed Medhat AliNo ratings yet
- H.B. Xiao, M. Krucker, K. Putzbach, K. Albert: A B A A ADocument9 pagesH.B. Xiao, M. Krucker, K. Putzbach, K. Albert: A B A A AJuan Camilo ZuluagaNo ratings yet
- Tujan ProjDocument4 pagesTujan ProjMayson BaliNo ratings yet
- Sensors & Actuators B. Chemical 304 (2020) 127396Document15 pagesSensors & Actuators B. Chemical 304 (2020) 127396ashwanirubalNo ratings yet
- Radhakrishnanand2008 PDFDocument5 pagesRadhakrishnanand2008 PDFnayelyNo ratings yet
- Development and Validation of Liquid Chromatography Method For Simultaneous Estimation of Miconazole and Clobetasol andDocument13 pagesDevelopment and Validation of Liquid Chromatography Method For Simultaneous Estimation of Miconazole and Clobetasol andvinayNo ratings yet
- Quantitative Determination of L-DOPA in PDFDocument6 pagesQuantitative Determination of L-DOPA in PDFMd.Mahfuzur RahmanNo ratings yet
- STD Method Clorofila A & BDocument20 pagesSTD Method Clorofila A & BDavid ZapataNo ratings yet
- The Strategy of Surfactant Analysis by HPLCDocument6 pagesThe Strategy of Surfactant Analysis by HPLCAlex100% (1)
- Validated Spectrophotometric Method For The Determination of Chloramphenicol in Pure and in Its Dosage FormDocument6 pagesValidated Spectrophotometric Method For The Determination of Chloramphenicol in Pure and in Its Dosage FormNin TiyasNo ratings yet
- Mitijps PaperDocument7 pagesMitijps PaperBrijeshkunvar MishraNo ratings yet
- Analyt10 Special KilicarslankucukDocument29 pagesAnalyt10 Special KilicarslankucukMaikel Perez NavarroNo ratings yet
- Journal of Pharmaceutical Analysis 2014Document7 pagesJournal of Pharmaceutical Analysis 2014Fernando OvallesNo ratings yet
- Analysis of Polar Agents by LC/MSDocument16 pagesAnalysis of Polar Agents by LC/MSkassim AliNo ratings yet
- Electrochemical Determination of Gatifloxacin, Moxifloxacin and Sparfloxacin Fluoroquinolonic Antibiotics On Glassy Carbon Electrode in Pharmaceutical FormulationsDocument4 pagesElectrochemical Determination of Gatifloxacin, Moxifloxacin and Sparfloxacin Fluoroquinolonic Antibiotics On Glassy Carbon Electrode in Pharmaceutical FormulationsEmilia Peralta MolinaNo ratings yet
- RP HPTLC Method For Determination of Voriconazole in - 2017 - Arabian Journal oDocument6 pagesRP HPTLC Method For Determination of Voriconazole in - 2017 - Arabian Journal olucian_lovNo ratings yet
- Thermo - Drug Abuse in UrineDocument7 pagesThermo - Drug Abuse in UrineYoosu NguyenNo ratings yet
- Nadifloxacin - HPTLC Stability Indicating PDFDocument8 pagesNadifloxacin - HPTLC Stability Indicating PDFNájla KassabNo ratings yet
- Validated Spectroscopic Method For Estimation of Naproxen From Tablet FormulationDocument3 pagesValidated Spectroscopic Method For Estimation of Naproxen From Tablet FormulationAdil AminNo ratings yet
- The Rapid Detection of Antioxidants From SafflowerDocument7 pagesThe Rapid Detection of Antioxidants From SafflowerRaquel Manzano DuránNo ratings yet
- On-Site Water Testing for AntibioticsDocument5 pagesOn-Site Water Testing for AntibioticsZulkarnain AjoiNo ratings yet
- QBD Approach in The Comparative Study of Azurin and Dabrafenib Anticancer Agents by UPLC Mass SeptrometryDocument3 pagesQBD Approach in The Comparative Study of Azurin and Dabrafenib Anticancer Agents by UPLC Mass SeptrometryInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Development and Validation of New ICP-OES Analytical Technique To Quantify The Contents of Copper, Magnesium & Zinc in "Escitalopram Oxalate"Document8 pagesDevelopment and Validation of New ICP-OES Analytical Technique To Quantify The Contents of Copper, Magnesium & Zinc in "Escitalopram Oxalate"Deepakrao Bornare PatilNo ratings yet
- High Performance Thin Layer Chromatographic Method With Densitometry Analysis For Determination of Rivaroxaban From Its Tablet Dosage FormDocument4 pagesHigh Performance Thin Layer Chromatographic Method With Densitometry Analysis For Determination of Rivaroxaban From Its Tablet Dosage FormPinak PatelNo ratings yet
- Advancing Healthcare Predictions: Harnessing Machine Learning for Accurate Health Index PrognosisDocument8 pagesAdvancing Healthcare Predictions: Harnessing Machine Learning for Accurate Health Index PrognosisInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Diabetic Retinopathy Stage Detection Using CNN and Inception V3Document9 pagesDiabetic Retinopathy Stage Detection Using CNN and Inception V3International Journal of Innovative Science and Research TechnologyNo ratings yet
- Comparatively Design and Analyze Elevated Rectangular Water Reservoir with and without Bracing for Different Stagging HeightDocument4 pagesComparatively Design and Analyze Elevated Rectangular Water Reservoir with and without Bracing for Different Stagging HeightInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Design, Development and Evaluation of Methi-Shikakai Herbal ShampooDocument8 pagesDesign, Development and Evaluation of Methi-Shikakai Herbal ShampooInternational Journal of Innovative Science and Research Technology100% (3)
- Terracing as an Old-Style Scheme of Soil Water Preservation in Djingliya-Mandara Mountains- CameroonDocument14 pagesTerracing as an Old-Style Scheme of Soil Water Preservation in Djingliya-Mandara Mountains- CameroonInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Cyberbullying: Legal and Ethical Implications, Challenges and Opportunities for Policy DevelopmentDocument7 pagesCyberbullying: Legal and Ethical Implications, Challenges and Opportunities for Policy DevelopmentInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- The Impact of Digital Marketing Dimensions on Customer SatisfactionDocument6 pagesThe Impact of Digital Marketing Dimensions on Customer SatisfactionInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- The Utilization of Date Palm (Phoenix dactylifera) Leaf Fiber as a Main Component in Making an Improvised Water FilterDocument11 pagesThe Utilization of Date Palm (Phoenix dactylifera) Leaf Fiber as a Main Component in Making an Improvised Water FilterInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Dense Wavelength Division Multiplexing (DWDM) in IT Networks: A Leap Beyond Synchronous Digital Hierarchy (SDH)Document2 pagesDense Wavelength Division Multiplexing (DWDM) in IT Networks: A Leap Beyond Synchronous Digital Hierarchy (SDH)International Journal of Innovative Science and Research TechnologyNo ratings yet
- Formulation and Evaluation of Poly Herbal Body ScrubDocument6 pagesFormulation and Evaluation of Poly Herbal Body ScrubInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Auto Encoder Driven Hybrid Pipelines for Image Deblurring using NAFNETDocument6 pagesAuto Encoder Driven Hybrid Pipelines for Image Deblurring using NAFNETInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Electro-Optics Properties of Intact Cocoa Beans based on Near Infrared TechnologyDocument7 pagesElectro-Optics Properties of Intact Cocoa Beans based on Near Infrared TechnologyInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Explorning the Role of Machine Learning in Enhancing Cloud SecurityDocument5 pagesExplorning the Role of Machine Learning in Enhancing Cloud SecurityInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- A Survey of the Plastic Waste used in Paving BlocksDocument4 pagesA Survey of the Plastic Waste used in Paving BlocksInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- A Review: Pink Eye Outbreak in IndiaDocument3 pagesA Review: Pink Eye Outbreak in IndiaInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Navigating Digitalization: AHP Insights for SMEs' Strategic TransformationDocument11 pagesNavigating Digitalization: AHP Insights for SMEs' Strategic TransformationInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Automatic Power Factor ControllerDocument4 pagesAutomatic Power Factor ControllerInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Hepatic Portovenous Gas in a Young MaleDocument2 pagesHepatic Portovenous Gas in a Young MaleInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Studying the Situation and Proposing Some Basic Solutions to Improve Psychological Harmony Between Managerial Staff and Students of Medical Universities in Hanoi AreaDocument5 pagesStudying the Situation and Proposing Some Basic Solutions to Improve Psychological Harmony Between Managerial Staff and Students of Medical Universities in Hanoi AreaInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Review of Biomechanics in Footwear Design and Development: An Exploration of Key Concepts and InnovationsDocument5 pagesReview of Biomechanics in Footwear Design and Development: An Exploration of Key Concepts and InnovationsInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- The Effect of Time Variables as Predictors of Senior Secondary School Students' Mathematical Performance Department of Mathematics Education Freetown PolytechnicDocument7 pagesThe Effect of Time Variables as Predictors of Senior Secondary School Students' Mathematical Performance Department of Mathematics Education Freetown PolytechnicInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Mobile Distractions among Adolescents: Impact on Learning in the Aftermath of COVID-19 in IndiaDocument2 pagesMobile Distractions among Adolescents: Impact on Learning in the Aftermath of COVID-19 in IndiaInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Enhancing the Strength of Concrete by Using Human Hairs as a FiberDocument3 pagesEnhancing the Strength of Concrete by Using Human Hairs as a FiberInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Drug Dosage Control System Using Reinforcement LearningDocument8 pagesDrug Dosage Control System Using Reinforcement LearningInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Securing Document Exchange with Blockchain Technology: A New Paradigm for Information SharingDocument4 pagesSecuring Document Exchange with Blockchain Technology: A New Paradigm for Information SharingInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Intelligent Engines: Revolutionizing Manufacturing and Supply Chains with AIDocument14 pagesIntelligent Engines: Revolutionizing Manufacturing and Supply Chains with AIInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Formation of New Technology in Automated Highway System in Peripheral HighwayDocument6 pagesFormation of New Technology in Automated Highway System in Peripheral HighwayInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Perceived Impact of Active Pedagogy in Medical Students' Learning at the Faculty of Medicine and Pharmacy of CasablancaDocument5 pagesPerceived Impact of Active Pedagogy in Medical Students' Learning at the Faculty of Medicine and Pharmacy of CasablancaInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Supply Chain 5.0: A Comprehensive Literature Review on Implications, Applications and ChallengesDocument11 pagesSupply Chain 5.0: A Comprehensive Literature Review on Implications, Applications and ChallengesInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- The Making of Self-Disposing Contactless Motion-Activated Trash Bin Using Ultrasonic SensorsDocument7 pagesThe Making of Self-Disposing Contactless Motion-Activated Trash Bin Using Ultrasonic SensorsInternational Journal of Innovative Science and Research TechnologyNo ratings yet
- Solids Level Measurement Application Guide en 78224 PDFDocument144 pagesSolids Level Measurement Application Guide en 78224 PDFwalcalNo ratings yet
- Investigating Population Growth SimulationDocument11 pagesInvestigating Population Growth Simulationapi-3823725640% (3)
- Copia de Tissue Response To Dental CariesDocument7 pagesCopia de Tissue Response To Dental Cariesjorefe12No ratings yet
- Right To HealthDocument9 pagesRight To HealthPriya SharmaNo ratings yet
- Grade Eleven Test 2019 Social StudiesDocument6 pagesGrade Eleven Test 2019 Social StudiesClair VickerieNo ratings yet
- Indonesia Organic Farming 2011 - IndonesiaDOCDocument18 pagesIndonesia Organic Farming 2011 - IndonesiaDOCJamal BakarNo ratings yet
- AZ ATTR Concept Test Clean SCREENERDocument9 pagesAZ ATTR Concept Test Clean SCREENEREdwin BennyNo ratings yet
- Myth and Realism in The Play A Long Day's Journey Into Night of Eugene O'neillDocument4 pagesMyth and Realism in The Play A Long Day's Journey Into Night of Eugene O'neillFaisal JahangeerNo ratings yet
- 2.1. Pharmacological Therapeutics. 2.2. Basic Cardiac Life Support (BCLS) and Advanced Cardiac Life Support (ACLS) in Neonates and ChildDocument3 pages2.1. Pharmacological Therapeutics. 2.2. Basic Cardiac Life Support (BCLS) and Advanced Cardiac Life Support (ACLS) in Neonates and Childclint xavier odangoNo ratings yet
- Manual Masina de Spalat Slim SamsungDocument1,020 pagesManual Masina de Spalat Slim SamsungPerfectreviewNo ratings yet
- Heat Exchanger Sodium SilicateDocument2 pagesHeat Exchanger Sodium SilicateChristopher BrownNo ratings yet
- Practical Examination Marking Guideline Grade 12 Physical Science 2019 PDFDocument5 pagesPractical Examination Marking Guideline Grade 12 Physical Science 2019 PDFWonder Bee Nzama100% (1)
- Grade 3 science syllabus 1st and 2nd semesterDocument2 pagesGrade 3 science syllabus 1st and 2nd semesterelyzabeth SibaraniNo ratings yet
- Synthesis, Experimental and Theoretical Characterizations of A NewDocument7 pagesSynthesis, Experimental and Theoretical Characterizations of A NewWail MadridNo ratings yet
- Chapter 4Document26 pagesChapter 4Lana AlakhrasNo ratings yet
- Journalize The Following Transactions in The Journal Page Below. Add Explanations For The Transactions and Leave A Space Between EachDocument3 pagesJournalize The Following Transactions in The Journal Page Below. Add Explanations For The Transactions and Leave A Space Between EachTurkan Amirova100% (1)
- TDS Versimax HD4 15W40Document1 pageTDS Versimax HD4 15W40Amaraa DNo ratings yet
- TSS-TS-TATA 2.95 D: For Field Service OnlyDocument2 pagesTSS-TS-TATA 2.95 D: For Field Service OnlyBest Auto TechNo ratings yet
- Annex 8 Qualification of BalancesDocument11 pagesAnnex 8 Qualification of BalancesMassimiliano PorcelliNo ratings yet
- Subaru Forester ManualsDocument636 pagesSubaru Forester ManualsMarko JakobovicNo ratings yet
- Family MedicineDocument156 pagesFamily MedicinedtriggNo ratings yet
- Business Startup Practical Plan PDFDocument70 pagesBusiness Startup Practical Plan PDFShaji Viswanathan. Mcom, MBA (U.K)No ratings yet
- Parasitology Lecture Hosts, Symbiosis & TransmissionDocument10 pagesParasitology Lecture Hosts, Symbiosis & TransmissionPatricia Ann JoseNo ratings yet
- Hotel Housekeeping EQUIPMENTDocument3 pagesHotel Housekeeping EQUIPMENTsamahjaafNo ratings yet
- Study On Marketing Strategies of Fast Food Joints in IndiaDocument35 pagesStudy On Marketing Strategies of Fast Food Joints in IndiaNiveditaParaashar100% (1)
- Exercise 4 Summary - KEY PDFDocument3 pagesExercise 4 Summary - KEY PDFFrida Olea100% (1)
- Quality Nutrition and Dietetics PracticeDocument3 pagesQuality Nutrition and Dietetics PracticeNurlienda HasanahNo ratings yet
- Funds Flow Statement ExplainedDocument76 pagesFunds Flow Statement Explainedthella deva prasad0% (1)
- Sub Erna RekhaDocument2 pagesSub Erna Rekhasurabhi mandalNo ratings yet
- Measure BlowingDocument52 pagesMeasure BlowingLos Ángeles Customs GarageNo ratings yet